 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cc02_g39470 | ||||||||
| Common Name | GSCOC_T00027710001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Rubiaceae; Ixoroideae; Coffeeae; Coffea
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 607aa MW: 66433.8 Da PI: 6.837 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.5 | 1e-12 | 439 | 485 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
Cc02_g39470 439 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAIAYITELQ 485
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 9.3E-56 | 52 | 240 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.85 | 435 | 484 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 3.79E-18 | 438 | 497 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.33E-14 | 438 | 489 | No hit | No description |
| Pfam | PF00010 | 3.4E-10 | 439 | 485 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.0E-18 | 439 | 498 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.3E-16 | 441 | 490 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 607 aa Download sequence Send to blast |
MKFEVGMGGL IWNDEDKTVV AAVLGTKAFD YLFSSSLSAE GSLMAMGNDE NLQNKLSDLV 60 EHPNSSNFSW NYAIFWQISR SKSGDLVLGW GDGCCREPRE GEESEVTRIL NLRLEDETQQ 120 RMRKRVLQKL HTCFGGSDED SYAFGLDRVT DTEMFFLASM YFSFPRGEGG PGKSFGSGKH 180 LWVSDALKSS ADYCVRSFLA KSAGMQTLVL IPTDVGVVEL GSVRSIPESL ELVQSARSSF 240 SSFSSFIRAK QAAAVVMGAD KKDGNAPCPN LGIGERPDGI PKIFGQDLNP GNPQSREKLA 300 IRKPNEEHWD GYPNGNRIAF PSPRNGIHGA TWTTFGHVKP GNGAELYSPQ TQTSKLQGLV 360 NGSGDEFRLG SFQHQKPTQM QIDFTGATSR PVISRPLCVD SEHSDVEASC KEDVSVLGDE 420 KRPRKRGRKP ANGREEPLNH VEAERQRREK LNQRFYALRA VVPNISKMDK ASLLGDAIAY 480 ITELQKKLKD MESERDKFGS ASRDALTSEA SPSSDVQSLV PEIDIQGAHD KVIVKVSCPL 540 DTHPASRVIH GLKEAGVNVI ESKLAAGNDK VFHTFVVKPQ GSEQLTKDKL IAAISRSCES 600 TSLQAL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 3e-28 | 433 | 495 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 3e-28 | 433 | 495 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 3e-28 | 433 | 495 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 3e-28 | 433 | 495 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 3e-28 | 433 | 495 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 3e-28 | 433 | 495 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 3e-28 | 433 | 495 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 3e-28 | 433 | 495 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 420 | 428 | KRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
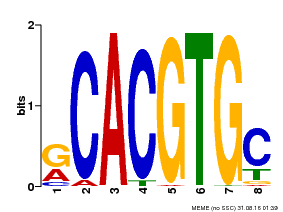
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027160117.1 | 0.0 | transcription factor bHLH13 isoform X1 | ||||
| Swissprot | A0A3Q7ELQ2 | 0.0 | MTB1_SOLLC; Transcription factor MTB1 | ||||
| TrEMBL | A0A068UK62 | 0.0 | A0A068UK62_COFCA; Uncharacterized protein | ||||
| STRING | XP_009781532.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3585 | 23 | 48 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 0.0 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



