 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cc02_g24540 | ||||||||
| Common Name | GSCOC_T00028871001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Rubiaceae; Ixoroideae; Coffeeae; Coffea
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 327aa MW: 35683.9 Da PI: 8.5816 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.9 | 2.2e-41 | 172 | 249 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cq+egC+adl++ak+yhrrhkvCe+hska++v++sgl+qrfCqqCsrfh lsefD++krsCr+rLa+hn+rrrk+q+
Cc02_g24540 172 RCQAEGCNADLTNAKHYHRRHKVCEFHSKAATVIASGLTQRFCQQCSRFHLLSEFDNGKRSCRKRLADHNRRRRKSQQ 249
6**************************************************************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 3.6E-35 | 167 | 234 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.547 | 170 | 247 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.96E-38 | 171 | 250 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 3.0E-31 | 173 | 246 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009554 | Biological Process | megasporogenesis | ||||
| GO:0009556 | Biological Process | microsporogenesis | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 327 aa Download sequence Send to blast |
MLDYEWGNPS AVMFTGDDST QDADQNRQLF DPYGTQNFGE ATAALVHQNA HYSAAAHHQH 60 TGNPFHHPHD PQGQSSNAHF SSLFDPRAAY GASSFNPHHQ ASMLSLEPAG STGFMVIPKS 120 EPVVGGADFT AAGRIGLNLG GRTYFSSSED DFVSRLYRRS RVVEPGSVNS PRCQAEGCNA 180 DLTNAKHYHR RHKVCEFHSK AATVIASGLT QRFCQQCSRF HLLSEFDNGK RSCRKRLADH 240 NRRRRKSQQQ QQQPNQEHCN TNSKKSPNDN TRSPPDSGAH SSSVTVAVSP PRISLDCFRQ 300 RASYQGGGAA ATNSSASTSS LFFSDG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-30 | 165 | 246 | 3 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. Binds specifically to the 5'-GTAC-3' core sequence. Involved in development and floral organogenesis. Required for ovule differentiation, pollen production, filament elongation, seed formation and siliques elongation. Also seems to play a role in the formation of trichomes on sepals. May positively modulate gibberellin (GA) signaling in flower. {ECO:0000269|PubMed:12671094, ECO:0000269|PubMed:16095614, ECO:0000269|PubMed:17093870}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00090 | SELEX | Transfer from AT1G02065 | Download |
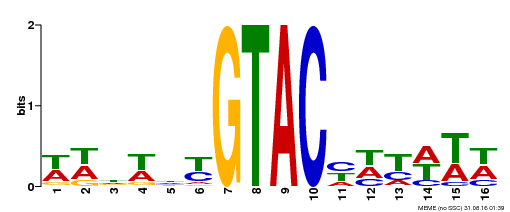
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027112970.1 | 0.0 | squamosa promoter-binding-like protein 8 | ||||
| Refseq | XP_027163563.1 | 0.0 | squamosa promoter-binding-like protein 8 | ||||
| Swissprot | Q8GXL3 | 1e-81 | SPL8_ARATH; Squamosa promoter-binding-like protein 8 | ||||
| TrEMBL | A0A068UMK9 | 0.0 | A0A068UMK9_COFCA; Uncharacterized protein | ||||
| STRING | Migut.B01572.1.p | 1e-137 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4846 | 23 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G02065.1 | 7e-83 | squamosa promoter binding protein-like 8 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



