 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cc02_g15780 | ||||||||
| Common Name | GSCOC_T00014039001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Rubiaceae; Ixoroideae; Coffeeae; Coffea
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 372aa MW: 42357.6 Da PI: 4.5978 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 110.9 | 8.9e-35 | 13 | 104 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkellekik 98
Fl+k+ye+++d+ ++ ++sws n++sf+v+++ ef++++Lp+yFkh+nf+SF+RQLn+YgF+k++ e+ weF+++ F +g+ +ll++i
Cc02_g15780 13 FLTKTYEMVDDPLTNAIVSWSCNNRSFIVWNPPEFSRDLLPQYFKHNNFSSFIRQLNTYGFRKIDPEQ---------WEFANDDFVRGQPHLLKNIY 100
9********************999****************************************9998.........******************** PP
XXXX CS
HSF_DNA-bind 99 rkks 102
r+k
Cc02_g15780 101 RRKP 104
*985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 9.3E-38 | 7 | 97 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 6.7E-61 | 9 | 102 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| SuperFamily | SSF46785 | 1.22E-34 | 10 | 102 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF00447 | 2.9E-31 | 13 | 102 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 3.1E-19 | 13 | 36 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 3.1E-19 | 51 | 63 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 52 | 76 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 3.1E-19 | 64 | 76 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 372 aa Download sequence Send to blast |
MDEPCSSNVL PPFLTKTYEM VDDPLTNAIV SWSCNNRSFI VWNPPEFSRD LLPQYFKHNN 60 FSSFIRQLNT YGFRKIDPEQ WEFANDDFVR GQPHLLKNIY RRKPVHSHSG QGAVESQLSE 120 SERRRYKDDI ERLKSDKEAL LFELQRYKQE EQRFELQVLG TTESVQLMEQ KQKNMMSFLS 180 KILQRPILAL ELMPEPHIHE RKRRIPGNSY LNEDISMANT QINSQILQKE QLDASSLLAF 240 NKDLLEELDS TLSFWENILI NVGQACDRLT SSPALVESTS CAGSPVVSYT EPNPKMSEID 300 MDCDQNTNTA VINDTANSDE LVAMNPTNVP TNGPTGVNDV FWEQFLTENP GSTNSSEVKV 360 NIMIMAGFGG T* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 6e-24 | 7 | 102 | 24 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 6e-24 | 7 | 102 | 24 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 6e-24 | 7 | 102 | 24 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
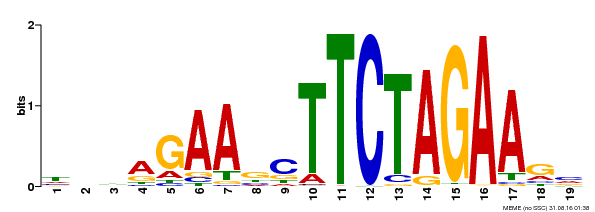
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027105613.1 | 0.0 | heat stress transcription factor A-4c-like | ||||
| Refseq | XP_027105614.1 | 0.0 | heat stress transcription factor A-4c-like | ||||
| TrEMBL | A0A068TLR4 | 0.0 | A0A068TLR4_COFCA; Uncharacterized protein | ||||
| STRING | XP_009796383.1 | 1e-141 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3448 | 23 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 7e-95 | heat shock transcription factor A4A | ||||



