 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10018473m | ||||||||
| Common Name | CICLE_v10018473mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1635aa MW: 184227 Da PI: 8.7 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.8 | 0.00036 | 1543 | 1565 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
Ciclev10018473m 1543 CPvkGCGKKFFSHKYLVQHRRVH 1565
9999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 11.7 | 0.00081 | 1601 | 1627 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
Ciclev10018473m 1601 YVCAepGCGQTFRFVSDFSRHKRKtgH 1627
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.7E-16 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.437 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 6.3E-15 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 4.9E-52 | 191 | 360 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 32.499 | 191 | 360 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.73E-27 | 209 | 381 | No hit | No description |
| Pfam | PF02373 | 4.1E-37 | 224 | 343 | IPR003347 | JmjC domain |
| SMART | SM00355 | 14 | 1518 | 1540 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0045 | 1541 | 1565 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.362 | 1541 | 1570 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 3.7E-5 | 1542 | 1564 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1543 | 1565 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.19E-9 | 1557 | 1599 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.8E-9 | 1565 | 1593 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0014 | 1571 | 1595 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.741 | 1571 | 1600 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1573 | 1595 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.25E-8 | 1589 | 1623 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.3E-9 | 1594 | 1624 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.468 | 1601 | 1632 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.62 | 1601 | 1627 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1603 | 1627 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1635 aa Download sequence Send to blast |
MAEPVQQQDI LPWLKTLPVA PEFHPTLAEF QDPIAYIFKI EKEASQYGIC KIVPPVPPPP 60 KKTAITFLNR SLAQRAAATG GATSSSGPTF TTRQQQIGFC PRKPRPVQKP VWQSGEYYTF 120 QEFETKAKNF EKSYLKKCGN KKAALSALEI ESLYWKASVD KPFSVEYAND MPGSAFVPVR 180 KIREAVGEGV TVGETPWNMR GVSRAKGSLL RFMKEEIPGV TSPMVYIAML FSWFAWHAED 240 HDLHSLNYLH MGASKTWYGV PMEAANAFEE VVRVHGYGEE INPLVTFATL GEKTTMISPE 300 VFVGAGVPCC RLVQNAGEFV VTFPRAYHMG FSHGFNCGEA ANIATPEWLN IAKDAAIRRA 360 SINYPPMVSH FQLLYDLAIA MHSSLPVAVS AKPRSSRLKD KNKDEGETLV KELFVQDVAQ 420 NNELLHVLGQ GSPIVLLPQS SSGALGANPW IPLGLCSYRE AIKSSGGLVS NDIMVGKNNG 480 INPVKGYCSV KGKFASLYAR NSSLSETDNI RNWNSQILST DTERQNTVQG DRSSDQRLFS 540 CVTCGILSFA CVAVIQPREP TARYLMSADC SFFNDWIVGS GVSGAFRAAG EDVIASEHNS 600 RSRWIGKSGR NSLYDVPVQS ANQIQAVDES NETISDRETK GDTSALNLLA ITYGNSSDSE 660 EEQVEPNVPM CDDKETKLTE CLLERKYQQN FHAAAAAAGS QDLSFISLDC EDEASLQISN 720 VQPEFRRDYL NDKNPQMSEC SVQFETDKHD CSKPNGFDGC FGDPIAASYA SKCAPVIHGG 780 ENVEFSKAIV PVMNAEMSFA PRSDEDSSRM HVFCLEHAVE VEQQLRPIGG VDIFLLCHPD 840 YPKMVAEAKL VAEELGIDSL CDEISFRVAT KEDEKRIHLS LDSEDAIPGN GDWAVKLGIN 900 LFYSANLSRS PLYSKQMPYN SIIYNAFGRS SPASSPNKYD NGRRPARQRK VVAGKWCGRV 960 WMSNQVHPFL VQKDPEEQEL ERSFHAWTTP DENFERKPES ICQTTSTLVT RKYSRKRKMV 1020 AESVSTKKAK CIDTEDAGSK YSLEGDTRIQ QRRILRNKPA KLMEKKDVDL PDSSEVSSYQ 1080 QKRSVSRRKQ AKCIQREVGD SNDALPGNSL IQYRRIPKRK HAKCIGREDA VLDDLTDDSS 1140 LKQYRRIPRS KLAKHVARED EVSDNSLRGT SDRQHTSIPK GKEFTCIDRD DAISDDSLQD 1200 NSRQLQFNRV PKSKQAEWIE REDAASDDSL EDYPHRLQHK KIPNIKQPKC IESEDPVLDN 1260 LLEKNTHRLH HQRILKSKRA KCVEREDAVS DDLLEDSSHM LPRRSQKSKQ AKWMREDAVS 1320 DDSLEDNCHQ QHKRIPRDKQ SKCTEMEDTV FYDSAEDKIR QFRRIARKRA NFTEREDAVL 1380 YDLLDNKSHR RHCRTLRSKQ LRTETLRKMK QETPSHMKPG KSRLTKQETS RLVKQVTSRQ 1440 HSVKSDQNAK LFDSVVEQEL EGGPSTRLRK RIPKPQKEFE TKPKEKNQAA KKKVKNASVV 1500 KAPAGLNNAK IKDEEAGYHC DMEGCTMSFG TKQELVLHKK NICPVKGCGK KFFSHKYLVQ 1560 HRRVHLDDRP LKCPWKGCKM TFKWAWARTE HIRVHTGARP YVCAEPGCGQ TFRFVSDFSR 1620 HKRKTGHSAK KSRG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 1e-74 | 7 | 384 | 2 | 356 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 1e-74 | 7 | 384 | 2 | 356 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1013 | 1018 | SRKRKM |
| 2 | 1113 | 1119 | RRIPKRK |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in the shoot apical meristem and primary and secondary root tips, and lower expression in cotyledons, leaves and root axis along vascular tissues. Detected in inflorescences, stems and siliques. {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18713399}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
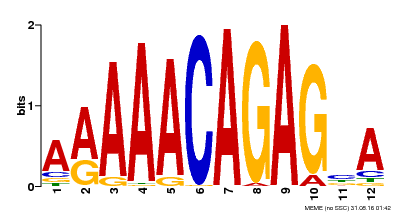
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024044095.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | V4TIR4 | 0.0 | V4TIR4_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006440007.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5259 | 28 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10018473m |
| Entrez Gene | 18048471 |



