 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10012759m | ||||||||
| Common Name | CICLE_v10012016mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 211aa MW: 24533 Da PI: 8.1384 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.3 | 0.0005 | 25 | 47 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y C +Cg + s ks+ +Hi tH
Ciclev10012759m 25 YYCDYCGICRSKKSLILSHIATH 47
89********************6 PP
| |||||||
| 2 | zf-C2H2 | 22.3 | 3.4e-07 | 68 | 89 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg sF+ + +Lk+H+ +H
Ciclev10012759m 68 TCEECGASFKKPAYLKQHMLSH 89
7*******************99 PP
| |||||||
| 3 | zf-C2H2 | 18.6 | 5.1e-06 | 95 | 119 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ dC+ s++rk++L+rH+ +H
Ciclev10012759m 95 FRCSvdDCHASYRRKDHLTRHLLRH 119
89*********************99 PP
| |||||||
| 4 | zf-C2H2 | 16.2 | 2.9e-05 | 124 | 149 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+kCp +C++ F+ + n+krH + H
Ciclev10012759m 124 FKCPieNCNREFTIQGNMKRHVKElH 149
89********************9986 PP
| |||||||
| 5 | zf-C2H2 | 19.2 | 3.4e-06 | 164 | 188 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
y C+ Cgk+F+ s L++H +H
Ciclev10012759m 164 YICQeiGCGKVFKYASKLRQHEDSH 188
89*******************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50157 | 8.933 | 25 | 52 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 21 | 25 | 47 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.528 | 67 | 94 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0088 | 67 | 89 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 3.1E-7 | 68 | 88 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.75E-10 | 68 | 119 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 69 | 89 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-11 | 89 | 121 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0066 | 95 | 119 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.827 | 95 | 124 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 97 | 119 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.19E-6 | 122 | 150 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-6 | 122 | 146 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.637 | 124 | 155 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0015 | 124 | 149 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 126 | 150 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 4.7E-8 | 156 | 188 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.58E-5 | 160 | 188 | No hit | No description |
| PROSITE profile | PS50157 | 10.803 | 164 | 188 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0033 | 164 | 188 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 166 | 188 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 211 aa Download sequence Send to blast |
MEEVSDMEEE KGLEERPKSR VVRRYYCDYC GICRSKKSLI LSHIATHHKE DEERVDGDGE 60 TKGVQCNTCE ECGASFKKPA YLKQHMLSHL LERPFRCSVD DCHASYRRKD HLTRHLLRHQ 120 GKLFKCPIEN CNREFTIQGN MKRHVKELHH EGTTSTDSGD QKQYICQEIG CGKVFKYASK 180 LRQHEDSHGL LLKFQINMKF CNIIFVIVNV * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 1e-16 | 71 | 207 | 19 | 148 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 1e-16 | 71 | 207 | 19 | 148 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Both isoforms 1 and 2 transcripts are expressed in seedlings and both TFIIIA and TFIIIA-C are present at similar ratios in 10-day-old seedlings (at protein level). In correlation with the reorganization of 5S rDNA chromatin to a mature state, full-length isoform 1 protein TFIIIA with transcriptional activity accumulates and permits de novo transcription of 5S rRNA. Isoforms 1 and 2 transcripts accumulate in various tissues of the reproductive phase, including flowers and siliques, but only isoform 2 is present in mature seeds. In flowers, both TFIIIA and TFIIIA-C are present at similar ratios (at protein level). Very low amounts of TFIIIA are found in extracts from fresh siliques compared to TFIIIA-C. The amount of full-length TFIIIA protein progressively increases from fresh siliques to seeds concomitant with lower proportions of the shorter TFIIIA-C protein (at protein level) thus leading to 5S rRNA accumulation in the seed. {ECO:0000269|PubMed:22353599}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in seedlings, flowers, siliques and seeds. {ECO:0000269|PubMed:22353599}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
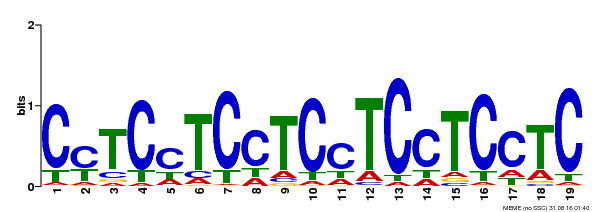
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006430925.1 | 1e-135 | transcription factor IIIA isoform X1 | ||||
| Swissprot | Q84MZ4 | 4e-54 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | V4T3S6 | 1e-154 | V4T3S6_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006430925.1 | 1e-134 | (Citrus clementina) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 2e-56 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10012759m |
| Entrez Gene | 18038914 |



