 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_41844 | ||||||||
| Common Name | KK1_043443 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 328aa MW: 36237.8 Da PI: 9.8271 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 44.1 | 4.6e-14 | 249 | 299 | 5 | 55 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkke 55
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++ N++L+ + e+ ++
C.cajan_41844 249 RRQRRMIKNRESAARSRARKQAYTFELEAEVAKLKELNRELQRKQAEIMEM 299
79************************************9998877777666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.6E-11 | 245 | 312 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.484 | 247 | 298 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 4.90E-27 | 249 | 303 | No hit | No description |
| Pfam | PF00170 | 6.9E-12 | 249 | 300 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 1.1E-13 | 249 | 298 | No hit | No description |
| SuperFamily | SSF57959 | 1.31E-9 | 249 | 297 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 252 | 267 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 328 aa Download sequence Send to blast |
MNFRNYGDDP TWDAMQGKTP TNVTLMRQPS VYSLTFDEFQ STMGGIGKDF GSMNMDELLK 60 NIWTAEETQA LGFSAGAGEG YNHNPISGGL QRQGSLTLPR TLNGSGNDGG SSMPQRQPTL 120 GEMTLEEFLV RAGVVREDVP QQIGKPNNNG SWFGDFHRPN TSTGLLLGFQ QPNRSNGNLG 180 LLGVVDHSMN VGMVSLPNAN VTATGASPSC KLSPNVITRS NGDTSSTSPV HYVINRGRKC 240 SAIEKVVERR QRRMIKNRES AARSRARKQA YTFELEAEVA KLKELNRELQ RKQAEIMEMQ 300 KDKPELDPAC RSRVTKIQCL RRTLTGPW |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Could participate in abscisic acid-regulated gene expression. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
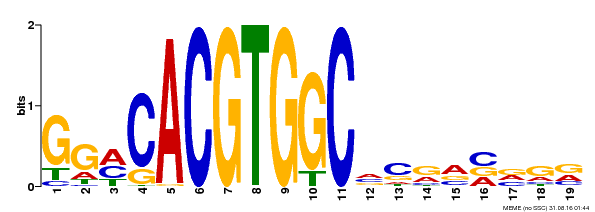
| Motif ID | Method | Source | Motif file |
| MP00087 | SELEX | Transfer from AT1G49720 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_41844 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by abscisic acid (ABA) and cold. {ECO:0000269|PubMed:10636868}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU418491 | 1e-109 | EU418491.1 Glycine max stress-related protein 1 (AREB1) mRNA, complete cds. | |||
| GenBank | KT031133 | 1e-109 | KT031133.1 Glycine max clone HN_CCL_105 BZIP transcription factor (Glyma02g14880.3) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020206247.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_029125682.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Swissprot | Q9M7Q5 | 5e-78 | AI5L4_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 4 | ||||
| TrEMBL | A0A151QYX3 | 0.0 | A0A151QYX3_CAJCA; ABSCISIC ACID-INSENSITIVE 5-like protein 4 | ||||
| STRING | XP_007156508.1 | 1e-160 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1962 | 34 | 81 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49720.1 | 2e-73 | abscisic acid responsive element-binding factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



