 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_31616 | ||||||||
| Common Name | KK1_031773 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1308aa MW: 146769 Da PI: 8.3797 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12 | 0.00065 | 1192 | 1217 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C C++sF +k +L +H r+ +
C.cajan_31616 1192 YQCDidGCTMSFGSKQELLHHKRNiC 1217
99********************9877 PP
| |||||||
| 2 | zf-C2H2 | 13.1 | 0.00029 | 1217 | 1239 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
C.cajan_31616 1217 CPvkGCGKKFFSHKYLVQHRRVH 1239
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 12 | 0.00063 | 1275 | 1301 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
C.cajan_31616 1275 YVCAepGCGQTFRFVSDFSRHKRKtgH 1301
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 4.4E-16 | 17 | 58 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.534 | 18 | 59 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 4.3E-14 | 19 | 52 | IPR003349 | JmjN domain |
| PROSITE profile | PS51184 | 33.68 | 181 | 357 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.48E-26 | 185 | 374 | No hit | No description |
| SMART | SM00558 | 8.5E-49 | 188 | 357 | IPR003347 | JmjC domain |
| Pfam | PF02373 | 1.8E-35 | 221 | 340 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 6.7E-4 | 1181 | 1214 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 6.9 | 1192 | 1214 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0045 | 1215 | 1239 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.944 | 1215 | 1244 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-6 | 1216 | 1243 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 3.47E-6 | 1216 | 1248 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 1217 | 1239 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.2E-10 | 1244 | 1269 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0014 | 1245 | 1269 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.614 | 1245 | 1274 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1247 | 1269 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.02E-10 | 1255 | 1299 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-10 | 1270 | 1298 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.62 | 1275 | 1301 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.468 | 1275 | 1306 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1277 | 1301 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1308 aa Download sequence Send to blast |
MGGASEDVLP WLKSLPVAPE YRPTVAEFQD PIAYIFKIEK EASKYGICKI IPPFPPSPKK 60 TAIANLNHSL AAVAGSTFTT RQQQIGFCPR RPRPVQRPVW QSGDHYTFHQ FESKAKSFEK 120 TYLKRHAKKG SGSASASASG SGLGLSPLET ETLFWKATLD KPFSVEYAND MPGSAFSPKC 180 RDAGDPSSLA DTPWNMRAVS RANGSLLRFM KEEIPGVTSP MVYVAMLFSW FAWHVEDHDL 240 HSLNYLHMGA GKTWYGVPRD AAVAFEEVVR VHGYGGEINP LVTFAILGEK TTVMSPEVFI 300 SAGVPCCRLV QNAGEFVVTF PRAYHTGFSH GFNCGEAANI ATPEWLRVAK DAAIRRASLN 360 YPPMVSHFQL LYDLALALCS RIPAGISAEP RSSRLKDKKK GEGETVIKEL FVQDVLQNND 420 LLHILGKGSP VVFLPRSSVD ISVCSKLRVG SQQSINVSNS EGMHSSKDNS KPIQRDNERE 480 TSQGEGLSDQ RLFSCVTCGI LSFSCVAIVQ PREPAARYLM SADCSFFNDW VVGSGVSSNK 540 FTIAHDRATI PEPNMYTGRM DEKDVPVQSS REALNTKSEN GNTALALLAS AYGNSSDSEE 600 DQIAIDDHSL KKQDYNITSG VTFENSRTVP NSTSNCSQDA HDAERSLSNK PMIPFDNKNA 660 SMVLQSDEDS SRMHVFCLEH AIEAEQQLRP IGGAHILLLC HPVMSVYVAK YKEIIFSSLM 720 ERLSNFDKNY PKIEAEAKLV AEDLGIDYMW KDTAYRHASK EDEETIQSAL DSEEAIPGNG 780 DWAVKLGINL FYSANLCRSP LYSKQMPYNS VIYYAFGCSS PASSPVEPKV YQRRVNKQKK 840 VVAGKWCGKV WMSNQVHPLL AKRDSEDVED EKLILGWILS DEKIERSEST PKSETTSRKS 900 GDYSPPYHHR KPISKRTNCT ESDAVSDDSL DDDDDYMQHR RNVNVEKAKF IDNDVVSNDS 960 VDCDSDWQQR AELSSKQVED TERDDISEDS LDVVSLQLQR KTSKGKHAKY ISEEDIISDD 1020 QMESHFQKRQ RRIPKSRQGR YITGKDITSD DQLELKVQKQ QRRNPKSRQA KYLNGEDIAS 1080 DDQLEGHFLK NQASRPLKRG SRILMKSKTP QQTKQSSHLR NKQSSNSQEF SLHMEEEEEG 1140 GPSTRLRKRA TKGQESEGKL KGKQTKRKKV KNATAAKVSV GHAKMKDGEP EYQCDIDGCT 1200 MSFGSKQELL HHKRNICPVK GCGKKFFSHK YLVQHRRVHE DERPLKCPWK GCKMTFKWAW 1260 ARTEHIRVHT GERPYVCAEP GCGQTFRFVS DFSRHKRKTG HSAKKSRQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 2e-79 | 7 | 378 | 4 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 2e-79 | 7 | 378 | 4 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1146 | 1168 | RKRATKGQESEGKLKGKQTKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
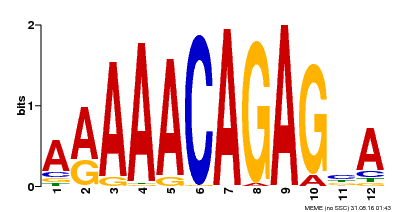
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_31616 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC235340 | 0.0 | AC235340.1 Glycine max strain Williams 82 clone GM_WBb0086A04, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007137965.1 | 0.0 | hypothetical protein PHAVU_009G169700g | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A151RVM6 | 0.0 | A0A151RVM6_CAJCA; Jumonji/ARID domain-containing protein 2 | ||||
| STRING | XP_007137965.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



