 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_28451 | ||||||||
| Common Name | KK1_027694 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 283aa MW: 32385.9 Da PI: 4.4898 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 63.4 | 3.2e-20 | 66 | 119 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++++q++ Le+ F +++++ e++++LAkklgL+ rqV vWFqNrRa++k
C.cajan_28451 66 KKHRLSSDQVHLLEKSFGEENKLEPERKTQLAKKLGLQPRQVAVWFQNRRARWK 119
67889************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 129 | 1.9e-41 | 65 | 157 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekk+rls++qv+lLE+sF ee+kLeperK++la++LglqprqvavWFqnrRAR+ktkqlE+dy+ Lk++yd+l ++ + + ke+e+L++e+++
C.cajan_28451 65 EKKHRLSSDQVHLLEKSFGEENKLEPERKTQLAKKLGLQPRQVAVWFQNRRARWKTKQLERDYDILKSSYDSLLSSYDLIAKENEKLKSEVES 157
69**************************************************************************************99875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 17.864 | 61 | 121 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.4E-18 | 63 | 125 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.67E-19 | 64 | 132 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.45E-17 | 66 | 122 | No hit | No description |
| Pfam | PF00046 | 1.7E-17 | 66 | 119 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 8.4E-22 | 67 | 128 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 4.3E-6 | 92 | 101 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 96 | 119 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 4.3E-6 | 101 | 117 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 4.3E-17 | 121 | 163 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001558 | Biological Process | regulation of cell growth | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 283 aa Download sequence Send to blast |
MESGRLFFDA SASRANNMLF LGTTELAFRA GRSMMSMEEA SKRRPFFTSP DELYDEEYYE 60 KQSPEKKHRL SSDQVHLLEK SFGEENKLEP ERKTQLAKKL GLQPRQVAVW FQNRRARWKT 120 KQLERDYDIL KSSYDSLLSS YDLIAKENEK LKSEVESLNE KLQLQAKEVP EEPLYDKKVD 180 PLPVDNIAPI FSTRVEDHLS SGSVGSAVVD EGSPQLVVDS VDSYFPADNY GGCVGPVERV 240 QSEEEDGSDD GRSYFDVLVV SETEHQNHEG EPLGWWSNMY YEG |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 113 | 121 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription activator involved in leaf development. Binds to the DNA sequence 5'-CAAT[AT]ATTG-3'. {ECO:0000269|PubMed:8535134}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00323 | DAP | Transfer from AT3G01470 | Download |
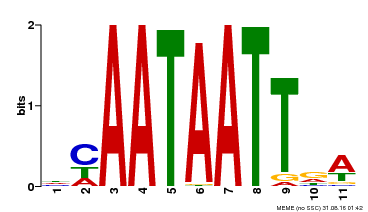
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_28451 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT093728 | 0.0 | BT093728.1 Soybean clone JCVI-FLGm-24I10 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020232165.1 | 0.0 | homeobox-leucine zipper protein HAT5 | ||||
| Swissprot | Q02283 | 3e-94 | HAT5_ARATH; Homeobox-leucine zipper protein HAT5 | ||||
| TrEMBL | A0A151S6Z4 | 0.0 | A0A151S6Z4_CAJCA; Homeobox-leucine zipper protein HAT5 | ||||
| STRING | GLYMA19G01300.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF11283 | 30 | 39 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 6e-82 | homeobox 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



