 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_27774 | ||||||||
| Common Name | KK1_028997 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1070aa MW: 120676 Da PI: 5.3301 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 179.9 | 3e-56 | 20 | 136 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrr 97
+e ++rwl++ ei++iL n++k++++ e+++ p+sgsl+L++rk++ryfrkDG++w+kkkdgktvrE+he+LK g+v+vl+cyYah+een++fqrr
C.cajan_27774 20 SEaQHRWLRPAEICEILSNHKKFQIAPEPAHMPSSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVREAHERLKAGSVDVLHCYYAHGEENENFQRR 115
4449******************************************************************************************** PP
CG-1 98 cywlLeeelekivlvhylevk 118
+yw+Lee+l++ivlvhy++vk
C.cajan_27774 116 TYWMLEEDLSHIVLVHYRQVK 136
******************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.567 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.1E-77 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.2E-49 | 21 | 134 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 1.56E-16 | 487 | 572 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 1.5E-6 | 487 | 564 | IPR002909 | IPT domain |
| Gene3D | G3DSA:2.60.40.10 | 2.1E-4 | 487 | 561 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF48403 | 3.58E-19 | 667 | 783 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.23E-15 | 669 | 781 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 2.2E-19 | 669 | 785 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 19.863 | 672 | 793 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 1.0E-8 | 694 | 789 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.0034 | 722 | 751 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.671 | 722 | 754 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 110 | 761 | 790 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 7.97E-7 | 891 | 944 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 12 | 894 | 916 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.026 | 896 | 915 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.455 | 896 | 924 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.16 | 917 | 939 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.584 | 918 | 942 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0032 | 920 | 937 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1070 aa Download sequence Send to blast |
MAETRHYVLP SQLDIEQIIS EAQHRWLRPA EICEILSNHK KFQIAPEPAH MPSSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVREAHER LKAGSVDVLH CYYAHGEENE NFQRRTYWML 120 EEDLSHIVLV HYRQVKGTKA NSTCAKENEE SLPYAQQTDK IMPKTEVDTS LSSTLRPHSY 180 QVPSQTIDTS MNSSQASEYE EAESAFNNHA SSDFYPFLKL QLPVEKIKAQ PADSYCPRPL 240 IDDQEKLPVI PGVNYISLTQ GNKMKDIHNF GLTYESSKPF GLSSWEGILE NNAESQHVPF 300 QPLFPETQPD NMGFNSNFSQ GDEIMVPHLT TSITEQHENG SLIQAEGNWQ AYGVDSLRMS 360 SWPIDSVYSG STCEVSCSNC DKEVDFQNSL EQYLEGNRNP DGMEDTYFTF KDGPPAEEGL 420 KKLDSFNQWM SKELGDVEES NKQSTSNAYW DTVESENEVG STTIPSQGHL DNFVLDPSIS 480 HDQLFSIIDY SPSWAFEESE VKVIISGRFL RSQHEAEQCK WSCMFGEVEV PAEIIANGVL 540 CCLTPPHKAG RVPFYVTCSN RLSCSEVREF DFQVNYTQEA NTEGENRGST FDTFSIRFGE 600 MLSLGHAFPQ NSDSISASEK FPLISKISSL LRGDDDDWDK LLKLTLEKDF SPENLQEQLL 660 ENLLKDKLHA WLIQKITEDG KGPNVLDEGG QGVLHFAAAL GYDWALEPTI VAGVNVNFRD 720 VNGWTALHWA AFCGRERTVA FLISLGAAPG ALTDPCPEHP SGRTPADLAS ANGHKGIAGY 780 LAESSLSAHL TTLDLNRDVG ENSGAKVVQR LQNTAQVHDL DGPSYELSLK DSLAAVCNAT 840 QAAARIHQVF RMQSFQRKQQ KEYGDDKFGI SDERALSLVT VNVKSHKSGP RDEPVHAAAI 900 RIQNKFRSWK GRREFLMIRQ RIVKIQAHVR GHQVRKSCGK IIWSVGILEK VILRWRRKGS 960 GLRGFKSEAI SEGTMIQDAS STEDDFDFLK EGRKQTEHRL QKALARVKSM VQYPEARDQY 1020 HRLLNVVTEI QENQVKHESS SMSSEEPRDF SNLNDLEALL DDDIFMPTAT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4hb5_A | 3e-12 | 661 | 810 | 13 | 157 | Engineered Protein |
| 4hb5_B | 3e-12 | 661 | 810 | 13 | 157 | Engineered Protein |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
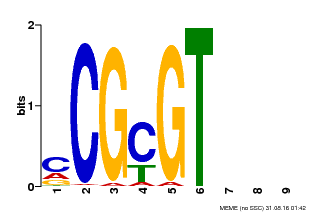
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_27774 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015034 | 0.0 | AP015034.1 Vigna angularis var. angularis DNA, chromosome 1, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020233265.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X1 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A151S369 | 0.0 | A0A151S369_CAJCA; Calmodulin-binding transcription activator 3 | ||||
| STRING | GLYMA08G14370.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8284 | 27 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



