 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_10906 | ||||||||
| Common Name | KK1_011226 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 576aa MW: 64006.4 Da PI: 6.7544 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 41.6 | 2.2e-13 | 405 | 451 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr+++P+ + K++Ka+ L A+ YI++Lq
C.cajan_10906 405 NHVEAERQRREKLNQRFYALRSVVPNiS------KMDKASLLGDAIAYINELQ 451
799***********************66......******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 2.0E-53 | 35 | 217 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| SuperFamily | SSF47459 | 1.57E-18 | 400 | 464 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.301 | 401 | 450 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.24E-14 | 404 | 455 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 3.7E-18 | 405 | 464 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 8.3E-11 | 405 | 451 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 6.7E-17 | 407 | 456 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 576 aa Download sequence Send to blast |
MVGAVLGGGA LEFLMRNSVW NESVLMAVGS EEGVQNNLSD LVERPNQSNF SWNYAIFWQL 60 SQSKSGDWVL CWGDGCCREP NEEEEVAVRG RGRLVEEEMH QRMRKRVLQK LHTTFGGSDE 120 DNYAVGLDHV TDTEMFFLAS MYFSFPRGHG GPGKCFASGN HLWLKSVSDY CVRSFLAKSA 180 GIQTVVLVPT DLGVVELGSV RILPESFQLL HTVKSLFSTH SSNALMRNSV WNEIVELGSV 240 RILPESFQLL HTVKSLFSTH SSNALFTGLA IGDGKVDAGV PIPKVFREKL AVRKMDDNNR 300 TWGGHPRNGL QAEIFAPRAS TSSVANAVRH DFRLGNTYQP QRQLQMQIDF TGATSRPSSV 360 KPVIGESELS DVEASCKEEQ PILADERKPR KRGRKPANGR DEPLNHVEAE RQRREKLNQR 420 FYALRSVVPN ISKMDKASLL GDAIAYINEL QAKVRIMEAE RERFGSTSKD ESVLEGNSSL 480 DIQDKEASRV NVQAVRDEVI VKVSCPFDAH PLSKVIQTLN EAQISVVESK LTAANDTIFH 540 TFVIKSQESE QLTKDKLIAV FSPTESNSLQ PLSSVG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 3e-29 | 399 | 461 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 3e-29 | 399 | 461 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 3e-29 | 399 | 461 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 3e-29 | 399 | 461 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 3e-29 | 399 | 461 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 3e-29 | 399 | 461 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 3e-29 | 399 | 461 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 3e-29 | 399 | 461 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 386 | 394 | RKPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
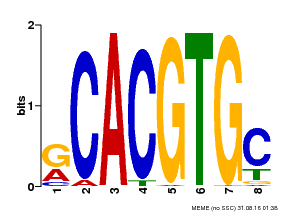
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_10906 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020212241.1 | 0.0 | transcription factor bHLH13 | ||||
| Swissprot | A0A3Q7ELQ2 | 0.0 | MTB1_SOLLC; Transcription factor MTB1 | ||||
| TrEMBL | A0A151TY93 | 0.0 | A0A151TY93_CAJCA; Transcription factor bHLH13 | ||||
| STRING | XP_007147321.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5448 | 32 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 1e-176 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



