 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa07g060910.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 341aa MW: 38614.4 Da PI: 6.4131 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 157.4 | 5.8e-49 | 14 | 140 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.kkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevls 94
l pGfrF+Ptd el+++yL++k++g+++++ vi+ev+iyk+ePwdLp ++ ++e+ew++F+ r +ky++g++++rat+ gyWkatgk+++v+s
Csa07g060910.1 14 LFPGFRFSPTDVELISYYLRRKIDGDEHSV-AVIAEVEIYKFEPWDLPeESKLKSENEWFYFCARGRKYPHGSQSRRATQLGYWKATGKERSVKS 107
579************************999.89***************433334678************************************** PP
NAM 95 kkgelvglkktLvfykgrapkgektdWvmheyrl 128
++++vg+k+tLvf+ grap+ge+t+W+mhey +
Csa07g060910.1 108 -GNQVVGTKRTLVFHIGRAPRGERTEWIMHEYCI 140
.999****************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.12E-54 | 6 | 155 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 52.419 | 14 | 156 | IPR003441 | NAC domain |
| Pfam | PF02365 | 2.7E-24 | 16 | 139 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009845 | Biological Process | seed germination | ||||
| GO:0009938 | Biological Process | negative regulation of gibberellic acid mediated signaling pathway | ||||
| GO:0033619 | Biological Process | membrane protein proteolysis | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0071472 | Biological Process | cellular response to salt stress | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 341 aa Download sequence Send to blast |
MSKEAEMSIA VSALFPGFRF SPTDVELISY YLRRKIDGDE HSVAVIAEVE IYKFEPWDLP 60 EESKLKSENE WFYFCARGRK YPHGSQSRRA TQLGYWKATG KERSVKSGNQ VVGTKRTLVF 120 HIGRAPRGER TEWIMHEYCI HGAPQDALVV CRLRKNADFR ASSSQRHIED GVAQDDGYVG 180 QSGGSEKEEK SYSVYDSELQ ISNGDIAESS NVVEDQAEIV DDCYAEILND DIIKLDEEAM 240 KASQASRPTN PTQQETIFSE ASSKRSKCGI KKESKETMNC YALFRIKNVP GTELSWRFPN 300 PFHIKKDDSQ RLMKNVLATT VFLAILVSFF WTILIARNSS * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 4e-43 | 4 | 138 | 10 | 138 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator activated by proteolytic cleavage through regulated intramembrane proteolysis (RIP), probably via metalloprotease activity. Regulates gibberellic acid-mediated salt-responsive repression of seed germination and flowering via FT, thus delaying seed germination under high salinity conditions. {ECO:0000269|PubMed:17410378, ECO:0000269|PubMed:18363782, ECO:0000269|PubMed:19704528, ECO:0000269|PubMed:19704545}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00280 | DAP | Transfer from AT2G27300 | Download |
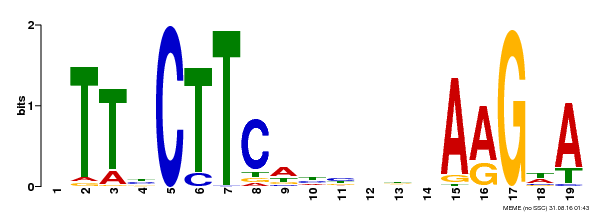
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa07g060910.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By high salt stress (PubMed:19704545, PubMed:17410378, PubMed:18363782, PubMed:19704528). Repressed by gibberellic acid (GA), but induced by the GA biosynthetic inhibitor paclabutrazol (PAC) (PubMed:18363782). Accumulates transiently in seeds upon imbibition (PubMed:19704545, PubMed:17410378, PubMed:18363782, PubMed:19704528). Induced by drought stress (PubMed:17158162). {ECO:0000269|PubMed:17158162, ECO:0000269|PubMed:17410378, ECO:0000269|PubMed:18363782, ECO:0000269|PubMed:19704528, ECO:0000269|PubMed:19704545}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT028934 | 0.0 | BT028934.1 Arabidopsis thaliana At2g27300 mRNA, complete cds. | |||
| GenBank | DQ056544 | 0.0 | DQ056544.1 Arabidopsis thaliana no apical meristem family protein (At2g27300) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010417881.1 | 0.0 | PREDICTED: NAC domain-containing protein 40 | ||||
| Swissprot | Q9XIN7 | 0.0 | NAC40_ARATH; NAC domain-containing protein 40 | ||||
| TrEMBL | R0HQ77 | 0.0 | R0HQ77_9BRAS; Uncharacterized protein | ||||
| STRING | XP_010417881.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10453 | 25 | 33 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G27300.1 | 0.0 | NTM1-like 8 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



