 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa06g022210.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1366aa MW: 153520 Da PI: 7.2923 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.5 | 0.00096 | 1248 | 1273 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C C++sF++ +L H r+ +
Csa06g022210.1 1248 YQCDmeGCTMSFTSEKQLALHKRNiC 1273
99********************9877 PP
| |||||||
| 2 | zf-C2H2 | 16.9 | 1.8e-05 | 1273 | 1295 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp CgksF ++ +L++H+r+H
Csa06g022210.1 1273 CPvrGCGKSFFSHKYLVQHQRVH 1295
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 13.6 | 0.00019 | 1331 | 1357 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C dCg++F+ s++ rH r+ H
Csa06g022210.1 1331 YVCAepDCGQTFRFVSDFSRHKRKtgH 1357
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 7.4E-17 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.813 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 5.7E-14 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 1.5E-53 | 200 | 369 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 34.854 | 203 | 369 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 2.31E-28 | 215 | 387 | No hit | No description |
| Pfam | PF02373 | 2.0E-38 | 233 | 352 | IPR003347 | JmjC domain |
| SMART | SM00355 | 8.5 | 1248 | 1270 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0018 | 1271 | 1295 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.422 | 1271 | 1300 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 6.0E-7 | 1272 | 1299 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1273 | 1295 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.96E-10 | 1287 | 1329 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 9.7E-9 | 1300 | 1324 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0015 | 1301 | 1325 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.928 | 1301 | 1330 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1303 | 1325 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 6.08E-8 | 1319 | 1353 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.0E-9 | 1325 | 1354 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 1.1 | 1331 | 1357 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.406 | 1331 | 1362 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1333 | 1357 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1366 aa Download sequence Send to blast |
MAVSEQSQDV FPWLKSLPVA PEFRPTLAEF QDPIAYIFKI EEEASKYGIC KILPPLPPPS 60 KKTSISNLNR SLAARAVGRV RDGGFGVCDY DGGPTFATRQ QQIGFCPRKQ RPVQRPVWQS 120 GEEYSFGEFE FKAKTFEKNY LKKCGKKSQL SPLEIETLYW RATVDKPFSV EYANDMPGSA 180 FIPLSLAAAR RRESGGDGGT VGETAWNMRA MARAEGSLLK FMKEDIPGVT SPMVYIAMMF 240 SWFAWHVEDH DLHSLNYLHM GASKTWYGVP KEAAPAFEEV VRVHGYGGEL NPLVTFSTLG 300 EKTTVMSPEV FVKAGIPCCR LVQNPGEFVV TFPGAYHSGF SHGFNFGEAS NIATPQWLTM 360 AKDAAIRRAA INYPPMVSHL QLLYDFALAL GSRVPTSIIA KPRSSRLKDK KRSEGERLTK 420 KLFVENIIHN NELLSSLGKG SPVGLLPQSS SDISVCSDLR IGSDLRTNQE KPIQLKSEEF 480 SSDSVMVGLS NDLRDAVSVK EKFTSLCERN RNHLASGEKE THETLSDAER REKGGAARLS 540 DQRLFSCVTC GVLSFDCVAI IQPKEAAAKY IMSADSSFFN DWTAASGSAN LGQAARSHYP 600 QSLEKHDVNY FSNVPVQTGH ERSSTTSLTN ADKDNGALGL LVSAYGDSSD SEEEDNKGLD 660 ASTSEGERKE NACAFQASSF DTDGNEVARD GRTSDFNSQR LTSKQNSLTK GDNSSLLELT 720 LPFVPRSDDD PCQLHVFCLE HAAEVEQQLR DIGGIHIMLL CHPEYPRIEA EAKIVAEELA 780 VNHEWNDTEF RNVTREDEET IQTALDNVEA KGGNSDWTVK LGINLSYSAI LSRSPLYSKQ 840 MPYNSVIYNV FGRSSPAVSS LSKPEVSGKR SFRQRKYVVG KWCGKVWMSH QVHPFLLEED 900 LEGEEERSCH LRTAMDEDVT GKRLFPCNVS RDAATMFGRK YCRKRKVRAK AVPRKKLTSF 960 KREDEVSDDT SEDHSYKQQW RASGKEEESY FETGHTASGD SSNQMSDQRK EIARHKDYNG 1020 IESDEEVSDR SLGEEYTVRE RAVSESSMEN GFQPYREKQS MYDRDEDDED DDMYKHPREI 1080 PRSKRTKVFR NPVSYNSEVA GVYQQRGRVS RSSRQANRLG GEYDSPENSF EELDLCSTGK 1140 RQTRSTAKRK AKTKTVQSPR DTKSRTLQEF ASGMKTEDLD SYMEGPSTRL RVRNQKPSRG 1200 SSESKPKKIG KKRSSNASFA RVASEEDVQE DVQEDEEAEN EEEECTGYQC DMEGCTMSFT 1260 SEKQLALHKR NICPVRGCGK SFFSHKYLVQ HQRVHSDDRP LKCPWKGCKM TFKWAWSRTE 1320 HIRVHTGARP YVCAEPDCGQ TFRFVSDFSR HKRKTGHSVK KTKKR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 1e-77 | 1245 | 1365 | 20 | 140 | Lysine-specific demethylase REF6 |
| 6a58_A | 1e-77 | 1245 | 1365 | 20 | 140 | Lysine-specific demethylase REF6 |
| 6a59_A | 1e-77 | 1245 | 1365 | 20 | 140 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 943 | 955 | KRKVRAKAVPRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
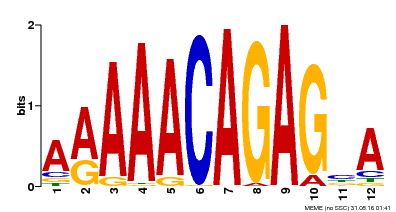
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa06g022210.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY664499 | 0.0 | AY664499.1 Arabidopsis thaliana relative of early flowering 6 (REF6) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010515159.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6-like | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | R0FN14 | 0.0 | R0FN14_9BRAS; Uncharacterized protein | ||||
| STRING | XP_010515159.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5259 | 28 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



