 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa03g001940.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 598aa MW: 66569.8 Da PI: 6.987 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.9 | 7.6e-13 | 440 | 485 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr+++P+ + K++Ka+ L Av YI++L
Csa03g001940.1 440 NHVEAERQRREKLNQRFYALRSVVPNiS------KMDKASLLGDAVSYINEL 485
799***********************66......***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.3E-51 | 49 | 237 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.326 | 436 | 485 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 8.51E-19 | 439 | 505 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 5.38E-14 | 439 | 490 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 3.5E-18 | 440 | 499 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 2.0E-10 | 440 | 485 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 4.8E-16 | 442 | 491 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 598 aa Download sequence Send to blast |
MNHIGGVVWN EDDKAIIASL LGKRALDYLL SNSVPNANLL MTVGSDDNLQ NKLSDLVERP 60 NSCNFSWNYA IFWQISRSKS GDLVLCWGDG SCREPKQGEK SDIVRILSMG REEETYQTMR 120 KRVLQKLHAM FGESEEDNCA LGLDRVTDTE MFLLSSMYFS FPRGEGGPGK CFASGKPLWL 180 SDVVNSGSDY CVRSFLAKSA GIQTIVLVPT DIGVVELGST SCLPESEESM LSIRSLFSST 240 LPPVRTLPAV ALPVVVAEKI DDNRTVTGSK IFGKDLHSSG FLHQHHQQQP HQQQQQQQQH 300 RQFREKLTVR KMDDRAPKRL DAYPNNNGNR FMFANPGSNN TLISPTWVQP ENYIRTNSVK 360 EVPSKDDFKF LPLQQASQRL LPPAQMQIDF SAASSRASEN NSDGEGGPEW ADTVGADESG 420 NNRPRKRGRR PANGRAEALN HVEAERQRRE KLNQRFYALR SVVPNISKMD KASLLGDAVS 480 YINELHAKLK VMEAEKERLG YSSNPPISLD SDINVQTSGE DVTVRINCPL ESHPASRIFH 540 AFEESKVEVI NSNLEVSQDT VLHTFVVKSQ ELTKEKLISA LSREQSSSVQ SRTSSGR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 2e-26 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 2e-26 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 2e-26 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 2e-26 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 2e-26 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 2e-26 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 2e-26 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 2e-26 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
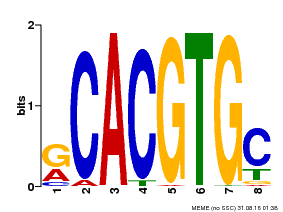
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa03g001940.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By UV treatment. {ECO:0000269|PubMed:12679534}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC023628 | 0.0 | AC023628.3 Arabidopsis thaliana chromosome I BAC F6F3 genomic sequence, complete sequence. | |||
| GenBank | AF251698 | 0.0 | AF251698.1 Arabidopsis thaliana putative transcription factor BHLH13 mRNA, complete cds. | |||
| GenBank | AY079012 | 0.0 | AY079012.1 Arabidopsis thaliana At1g01260/F6F3_25 mRNA, complete cds. | |||
| GenBank | AY120752 | 0.0 | AY120752.1 Arabidopsis thaliana transcription factor MYC7E, putative (At1g01260) mRNA, complete cds. | |||
| GenBank | BT004517 | 0.0 | BT004517.1 Arabidopsis thaliana At1g01260/F6F3_25 gene, complete cds. | |||
| GenBank | CP002684 | 0.0 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010480459.1 | 0.0 | PREDICTED: transcription factor bHLH13-like | ||||
| Swissprot | Q9LNJ5 | 0.0 | BH013_ARATH; Transcription factor bHLH13 | ||||
| TrEMBL | R0IMU8 | 0.0 | R0IMU8_9BRAS; Uncharacterized protein | ||||
| STRING | XP_010480459.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3758 | 27 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 0.0 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



