 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004504077.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1102aa MW: 126021 Da PI: 5.823 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 180.6 | 1.9e-56 | 19 | 135 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqr 96
+e ++rwl+++ei++iL+n++++++++++ + p+sgs++L++rk++ryfrkDG++w+kkkdgktvrE+he+LK g+v+vl+cyYah+e+n++fqr
XP_004504077.1 19 SEaQHRWLRSTEICQILTNYNSFQIASQPSHMPPSGSVFLFDRKVMRYFRKDGHNWRKKKDGKTVREAHERLKAGSVDVLHCYYAHGEQNENFQR 113
4449******************************************************************************************* PP
CG-1 97 rcywlLeeelekivlvhylevk 118
r+yw+Leeel++ivlvhy++vk
XP_004504077.1 114 RTYWMLEEELSHIVLVHYRQVK 135
*******************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 78.851 | 14 | 140 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 5.2E-76 | 17 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.0E-49 | 20 | 133 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 3.11E-16 | 513 | 598 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 6.2E-6 | 513 | 593 | IPR002909 | IPT domain |
| Gene3D | G3DSA:1.25.40.20 | 5.4E-19 | 700 | 815 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 18.404 | 703 | 824 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.18E-18 | 709 | 816 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 7.44E-14 | 709 | 812 | No hit | No description |
| Pfam | PF12796 | 4.0E-8 | 725 | 815 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.324 | 753 | 785 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.029 | 753 | 782 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 130 | 792 | 821 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 7.17E-7 | 920 | 975 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 34 | 925 | 947 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 6.888 | 927 | 955 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.22 | 929 | 946 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0032 | 948 | 970 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.901 | 949 | 973 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.3E-4 | 950 | 969 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1102 aa Download sequence Send to blast |
MAHNHLHPNQ FDTIEQILSE AQHRWLRSTE ICQILTNYNS FQIASQPSHM PPSGSVFLFD 60 RKVMRYFRKD GHNWRKKKDG KTVREAHERL KAGSVDVLHC YYAHGEQNEN FQRRTYWMLE 120 EELSHIVLVH YRQVKGVTKA NFICGKENEE YHPYAQQTDK VMPNTKMETF LSSSLNPLSY 180 QLQSQTMDTS INSFQASEYE EAESAFNSHE NSDLYSFLEL QHPFVHKIKA QLADSNSPLP 240 LKDDQERLPV IPQVDYISLS QANETKYINN ARLTCESSKL LGFSSWEDIL ENNAGCHNVI 300 SQPSFPETQH NNMNLNSTYQ GYDIMGQHFT ISITKQHENG SLIQAEGNWQ ASHFNSLSSS 360 NWPEDSACSG STCEVGYSDC EQEVNEVDLQ QSLEQFLLHP HQQHEVLMQN SPREILLNEE 420 DKLESELEVD RSIDGIEDTH FTSKKTLLDV SVAEEGLKKL DSFNQWMSKE LGDVEESSNR 480 STSSTYWDTV ESENEVGNYV LDPSISHDQL FSIIDYSPSW TFEYSEIKVL ISGRFLKSQH 540 EAEDCKWSCM FGEVEVPAEV IGNGVLCCHT PPHKAGRVPF YVTCSNRLAC SELREFDFCV 600 NYTQEVYTAG ENRSSITFDS FNKRFGDLLS LEHDFNHSLD SISVSEKSNE KYQLRSKISS 660 LLRREDDEWD KLLKFTLEKD FSPELVQEQL LEDLLKDKLH SWLLQKTTED GKGPNVLDES 720 GQGVLHFAAA LGYGWALEPT IIAGVNVNFR DVNGWTALHW AAVCGRERTV ASLISLGAAP 780 GALTDPCPKH PSGRTPADLA SENGHKGIAA YLAEYFLSAQ LKSLDLKRDL GENFGEKIIQ 840 RIQEQNTAKE VLSHELSLKD SLAAVCNATQ AAARIHQVFR VQSFQRKQKQ QKEYGDYKFG 900 VSDERALSLI TINAKSHKFG QCYEPVHIAA TRIQNKFRSW KGRKDFLIIR RRIVKIQAHV 960 RGHQVRKNYG KIVWSVGIME KVILRWRRKG SGLRGFKSEA ISDGTMVLGV SSSTEDDYDF 1020 LKEGRKQTEK RLEKALARVK SMAQYPDARD QYHRLLNVVT EIQENQVKQD WNFNNSEETR 1080 HVDNLVDLES LLNEDTFMPV DT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4hb5_A | 1e-11 | 709 | 845 | 25 | 158 | Engineered Protein |
| 4hb5_B | 1e-11 | 709 | 845 | 25 | 158 | Engineered Protein |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
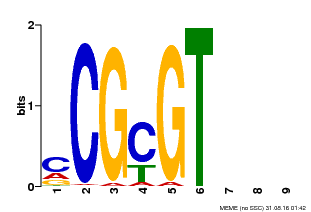
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004504077.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004504077.1 | 0.0 | calmodulin-binding transcription activator 1 | ||||
| TrEMBL | A0A1S2YFK3 | 0.0 | A0A1S2YFK3_CICAR; calmodulin-binding transcription activator 1 | ||||
| STRING | XP_004504077.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8284 | 27 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



