 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004496472.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 578aa MW: 66180.5 Da PI: 6.468 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 95.2 | 6e-30 | 49 | 133 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
rW++qe+laL+++r++m+ +r+++lk+plWeevs+k++e g++r++k+Ckek+en+ k++k++keg+++++se +t+++fdql+
XP_004496472.1 49 RWPRQETLALLKIRSDMDGVFRDSSLKGPLWEEVSRKLAELGYHRNAKKCKEKFENVYKYHKRTKEGKSGKKSEG-KTYRFFDQLQ 133
8********************************************************************985555.69******98 PP
| |||||||
| 2 | trihelix | 101.5 | 6.9e-32 | 389 | 474 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+k+ev+aLi++r+++e +++++ k+plWe++s+ m++ g++r++k+Ckekwen+nk+ykk+ke++k+r + s+tcpyf++lea
XP_004496472.1 389 RWPKSEVHALIRIRTSLEPKYQENGPKAPLWEDISAGMQRLGYNRNSKRCKEKWENINKYYKKMKESNKQR-RDGSKTCPYFNELEA 474
8*********************************************************************8.67778*******985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00717 | 0.1 | 46 | 108 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS50090 | 6.737 | 48 | 106 | IPR017877 | Myb-like domain |
| Pfam | PF13837 | 1.5E-19 | 48 | 135 | No hit | No description |
| CDD | cd12203 | 1.82E-22 | 48 | 113 | No hit | No description |
| SMART | SM00717 | 1.2 | 386 | 448 | IPR001005 | SANT/Myb domain |
| Pfam | PF13837 | 2.4E-22 | 388 | 475 | No hit | No description |
| PROSITE profile | PS50090 | 6.946 | 388 | 446 | IPR017877 | Myb-like domain |
| CDD | cd12203 | 1.68E-25 | 388 | 453 | No hit | No description |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 578 aa Download sequence Send to blast |
MELGDATVMD SSSGEIVAPE VHDGGDEKGR NGGEEDDKMM NITGSGGNRW PRQETLALLK 60 IRSDMDGVFR DSSLKGPLWE EVSRKLAELG YHRNAKKCKE KFENVYKYHK RTKEGKSGKK 120 SEGKTYRFFD QLQALEKQFS LSSYPPTSKP QPNNNIVSLP TKPNNTTTIS HVPSTNPTTL 180 ISPSPPPPLP PPTNATTTPT LTNNKNNNNV QYSLPNMNLF STTTTSTSSS TASDEDLEEK 240 YRKKRKWKDY FRRLTREVLI KQEEMQKKFL EAIDKREREH MAQQDAWRVQ EMNRINKEHE 300 LLVQERSTTA AKNAAVIAFL QKLSGQQNST IQDNFIQPPP PPQPTPPESA QTPISQLQIQ 360 PHEPVTSNNN IVEIHQNNGH KSGGGASSRW PKSEVHALIR IRTSLEPKYQ ENGPKAPLWE 420 DISAGMQRLG YNRNSKRCKE KWENINKYYK KMKESNKQRR DGSKTCPYFN ELEALYKEKN 480 KTFHNNIIKS SEMMEPLMVQ PEQQWRPPTT ITTTQYDEEE KKKKNSNEVE AKEEDYDDND 540 DDEYDEGDSM EEEEEDEGGV NSRYEVMTNK LSPVDTVE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds specific DNA sequence. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00244 | DAP | Transfer from AT1G76890 | Download |
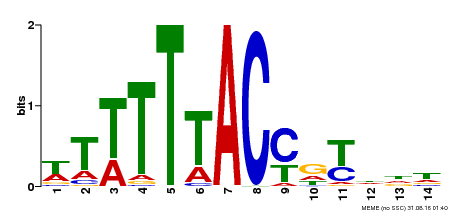
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004496472.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC225546 | 1e-114 | AC225546.4 Medicago truncatula clone mth2-78i17, complete sequence. | |||
| GenBank | BT149534 | 1e-114 | BT149534.1 Medicago truncatula clone JCVI-FLMt-19J11 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004496472.1 | 0.0 | trihelix transcription factor GT-2-like | ||||
| Swissprot | Q39117 | 1e-139 | TGT2_ARATH; Trihelix transcription factor GT-2 | ||||
| TrEMBL | A0A1S2XYC3 | 0.0 | A0A1S2XYC3_CICAR; trihelix transcription factor GT-2-like | ||||
| STRING | XP_004496472.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF373 | 34 | 181 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G76880.1 | 5e-79 | Trihelix family protein | ||||



