 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CA05g11860 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Capsiceae; Capsicum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 593aa MW: 65623.2 Da PI: 7.7408 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 35.7 | 1.5e-11 | 435 | 481 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ +I ++q
CA05g11860 435 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAIAHITDMQ 481
799***********************66......***************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.0E-53 | 45 | 233 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.056 | 431 | 480 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.83E-17 | 434 | 504 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 7.68E-13 | 434 | 485 | No hit | No description |
| Pfam | PF00010 | 4.4E-9 | 435 | 481 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.5E-16 | 435 | 494 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.3E-15 | 437 | 486 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 593 aa Download sequence Send to blast |
MTVLWSDEDK TMVAAVLGAK AFDYLMSSLV SAECSLISMG SNENLQNILS DLVERPNGSN 60 FSWNYAIFWQ ISRSKLGELV LGWGDGCCRE AREGEESEVT GILNLHLEDE AQQRMRKRVL 120 QKLHTFFGGT DEENYLFGLD RVTDTEMFFL ASMYFSFPHG QGGPGKCFGA GKHVWLSDVM 180 KSSVDYCSRS FLMKSAGMQT IVLIPTDVGV VELGSVRAIS ESLELVKSIK SCFSSFSSLG 240 RAKQAASVTV VAEKKDGNSS MFSSSFLRDQ SNGNPKIFGQ NLNSGCAQFR EKLLVRKPVD 300 RPLETYRNGN KGPFVNAQNG VCPVSWASFG NVKPVNSAEL YSPQAPTNNL RNFAVNGARE 360 EFRLNNLQHQ KHQKPGGMQI DFTNSRPVVY PVHTVESEHS DVEVSCKENH AGPADETRPR 420 KRGRKPANGR EEPLNHVEAE RQRREKLNQR FYALRAVVPN ISKMDKASLL GDAIAHITDM 480 QKKIRDMESE RGRLGSPNSE TQTRVADINI EAAGDEVIVR VKCPLEIHPL SKVIQVFKEA 540 QANVVESKVA AGNDTVYHTF VVKSSGSEQL TKEKLMAAFC GESNSLQPHS HQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 2e-28 | 429 | 506 | 2 | 79 | Transcription factor MYC2 |
| 5gnj_B | 2e-28 | 429 | 506 | 2 | 79 | Transcription factor MYC2 |
| 5gnj_E | 2e-28 | 429 | 506 | 2 | 79 | Transcription factor MYC2 |
| 5gnj_F | 2e-28 | 429 | 506 | 2 | 79 | Transcription factor MYC2 |
| 5gnj_G | 2e-28 | 429 | 506 | 2 | 79 | Transcription factor MYC2 |
| 5gnj_I | 2e-28 | 429 | 506 | 2 | 79 | Transcription factor MYC2 |
| 5gnj_M | 2e-28 | 429 | 506 | 2 | 79 | Transcription factor MYC2 |
| 5gnj_N | 2e-28 | 429 | 506 | 2 | 79 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
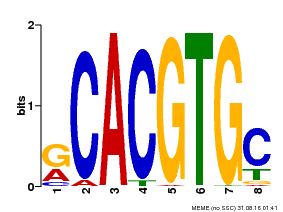
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975517 | 0.0 | HG975517.1 Solanum lycopersicum chromosome ch05, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016554600.1 | 0.0 | PREDICTED: transcription factor bHLH13-like isoform X1 | ||||
| Swissprot | A0A3Q7HES4 | 0.0 | MTB2_SOLLC; Transcription factor MTB2 | ||||
| TrEMBL | A0A1U8FFS3 | 0.0 | A0A1U8FFS3_CAPAN; Transcription factor bHLH13 | ||||
| TrEMBL | A0A2G2ZHT5 | 0.0 | A0A2G2ZHT5_CAPAN; transcription factor bHLH13-like isoform X1 | ||||
| STRING | XP_009627739.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3585 | 23 | 48 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 0.0 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



