 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CA02g29480 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Capsiceae; Capsicum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 288aa MW: 32422.6 Da PI: 4.5643 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 64.6 | 1.4e-20 | 66 | 119 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++++eq++ Le+ Fe ++++ e++++LA+klgL+ rqV vWFqNrRa++k
CA02g29480 66 KKRRLSPEQVHLLEKSFEAENKLEPERKTQLAQKLGLQPRQVAVWFQNRRARWK 119
56789************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 135.3 | 2.1e-43 | 65 | 156 | 1 | 92 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelk 92
ekkrrls eqv+lLE+sFe+e+kLeperK++la++LglqprqvavWFqnrRAR+ktkqlE+dy++Lk++yd+l ++ ++++k++++L++e+
CA02g29480 65 EKKRRLSPEQVHLLEKSFEAENKLEPERKTQLAQKLGLQPRQVAVWFQNRRARWKTKQLERDYDQLKSSYDSLLSDFDSIRKDNDQLKSEIV 156
69**************************************************************************************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.76E-20 | 53 | 123 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 18.285 | 61 | 121 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.1E-20 | 64 | 125 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 9.2E-18 | 66 | 119 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 8.91E-20 | 66 | 122 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 7.3E-22 | 67 | 128 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 4.4E-6 | 92 | 101 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 96 | 119 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 4.4E-6 | 101 | 117 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.6E-17 | 121 | 163 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001558 | Biological Process | regulation of cell growth | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 288 aa Download sequence Send to blast |
MGSGHIFFDP SSCHGNMLFL GNGDPLFRGP RSSMLKMEDS SKRRPFFSSP EELYDEEYYD 60 EQSPEKKRRL SPEQVHLLEK SFEAENKLEP ERKTQLAQKL GLQPRQVAVW FQNRRARWKT 120 KQLERDYDQL KSSYDSLLSD FDSIRKDNDQ LKSEIVSLME KLQGKVVGGS VVNEKSEALE 180 VDALTTLQVK VKTEDRLSSG SGGSAVVDEH SPQLVDSGDS YFHSTIDHEE YPGPGGCNIP 240 PPVDGLQSEE DDGSDDGSCH GYFSNAFVAS EEHHHEEGEE PLGWFWS* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 113 | 121 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
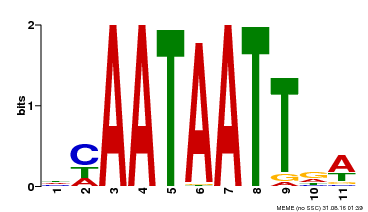
| UniProt | Probable transcription activator involved in leaf development. Binds to the DNA sequence 5'-CAAT[AT]ATTG-3'. {ECO:0000269|PubMed:8535134}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00323 | DAP | Transfer from AT3G01470 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM902288 | 0.0 | AM902288.1 Solanum lycopersicum mRNA for homeodomain leucine zipper protein (hb-1 gene). | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016561539.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein HAT5 | ||||
| Swissprot | Q02283 | 1e-94 | HAT5_ARATH; Homeobox-leucine zipper protein HAT5 | ||||
| TrEMBL | A0A1U8G0N0 | 0.0 | A0A1U8G0N0_CAPAN; Homeobox-leucine zipper protein HOX5 | ||||
| TrEMBL | A0A2G3AC40 | 0.0 | A0A2G3AC40_CAPAN; homeobox-leucine zipper protein HAT5 | ||||
| STRING | PGSC0003DMT400001322 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA7891 | 23 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 2e-80 | homeobox 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



