 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bv1_019790_iszi.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Betoideae; Beta
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 263aa MW: 29815.7 Da PI: 7.6841 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 93.8 | 7.7e-30 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
krien++ rqvtfskRr+g+lKKA+ELSvLCdae+a+i+fss+gklye+ss
Bv1_019790_iszi.t1 9 KRIENNISRQVTFSKRRTGLLKKAKELSVLCDAEIALIVFSSRGKLYEFSS 59
79***********************************************96 PP
| |||||||
| 2 | K-box | 87.1 | 3.4e-29 | 87 | 176 | 8 | 97 |
K-box 8 sleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkk 97
++ ea+++s+ qe+a L+++ e+L+r qRh+lGedL+sLslkeLq+Le+qLe++l + R++K +l+ eq++ l++ e++l + nk+L+ +
Bv1_019790_iszi.t1 87 HAIEAQTQSWLQEIAHLESQCESLKRAQRHMLGEDLGSLSLKELQHLEKQLESALAQTRARKAQLMAEQMDALRRSERKLGDLNKELKLR 176
367889********************************************************************************9865 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 32.744 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 3.4E-39 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 2.62E-31 | 2 | 83 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 3.30E-41 | 2 | 73 | No hit | No description |
| PRINTS | PR00404 | 8.5E-31 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 2.3E-25 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 8.5E-31 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 8.5E-31 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 5.0E-26 | 91 | 176 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 15.191 | 93 | 183 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0048437 | Biological Process | floral organ development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 263 aa Download sequence Send to blast |
MGRGKVELKR IENNISRQVT FSKRRTGLLK KAKELSVLCD AEIALIVFSS RGKLYEFSSI 60 ECLPKTIERY QRCGNNIELH HHNHQDHAIE AQTQSWLQEI AHLESQCESL KRAQRHMLGE 120 DLGSLSLKEL QHLEKQLESA LAQTRARKAQ LMAEQMDALR RSERKLGDLN KELKLRVGDQ 180 GHGFRPILDS NGCIPFIHQS QSTSFEYDPV TLQIGSYKPE PVVLEIGYAP HYPSHEDEGP 240 SNGNQRAGLG SDSTNFHQAG WLL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5f28_A | 4e-18 | 1 | 70 | 1 | 69 | MEF2C |
| 5f28_B | 4e-18 | 1 | 70 | 1 | 69 | MEF2C |
| 5f28_C | 4e-18 | 1 | 70 | 1 | 69 | MEF2C |
| 5f28_D | 4e-18 | 1 | 70 | 1 | 69 | MEF2C |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in flower development. {ECO:0000250|UniProtKB:Q0HA25}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00315 | DAP | Transfer from AT2G45650 | Download |
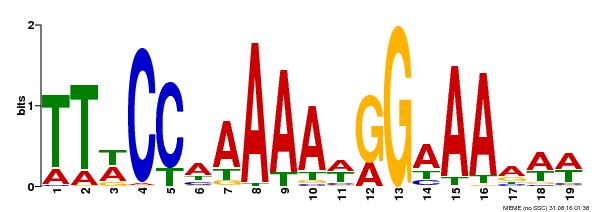
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010692324.1 | 0.0 | PREDICTED: truncated transcription factor CAULIFLOWER A isoform X1 | ||||
| Swissprot | Q8LLR1 | 2e-84 | MADS3_VITVI; Agamous-like MADS-box protein MADS3 | ||||
| TrEMBL | A0A0K9QM32 | 1e-162 | A0A0K9QM32_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010692324.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45650.1 | 3e-70 | AGAMOUS-like 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



