 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Brast03G018100.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 713aa MW: 77295.8 Da PI: 6.3527 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.9 | 3.2e-15 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+ E++++++a k++G W +I +++g ++t+ q++s+ qk+
Brast03G018100.1.p 24 RERWTEAEHKRFLEALKLYGRA-WQRIEEHVG-TKTAVQIRSHAQKF 68
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.84E-16 | 18 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.147 | 19 | 73 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.6E-16 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 6.5E-12 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.4E-12 | 24 | 67 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.8E-8 | 24 | 64 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.02E-8 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043433 | Biological Process | negative regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 713 aa Download sequence Send to blast |
MEMNSSGEET VIKVRKPYTI TKQRERWTEA EHKRFLEALK LYGRAWQRIE EHVGTKTAVQ 60 IRSHAQKFFT KLEKEAINNG TSPGEAHDID IPPPRPKRKP NCPYPRKGCL SSDTPTREVP 120 NDKSTKSNIS LTNSNAQTAG NAALQKLQGK ESSSCSEVLN LFRDAPSASF SSVNKSSSNH 180 GALMEIEATK TEIKDMTTMD GTSVSIDMEK DVKDINDQEM ERLNRIHMSA KYDHPHDDCL 240 DNSRQHVKLK PKSVEITYTD QHPARASHNQ MDRTGATGVP VIGTEGSPAD QTNDQGGVAK 300 GNMNCLHPTL AVDPKFDSSS TAQPITHNYA AFAPMMQCQC NQDAYKSSVN MSSTFSSMLV 360 STLLSNPAVH AAARLAASYW PSADTPVDPN QESPDDNAQG RHIGSPPSMA SIVAATVAAA 420 SAWWATQGLL PLFAPPMAFP FVPVPSASFP IADVQRASEN CPVDCTPKEC QVAQKQNKPE 480 AVVVVASSQS DESGNGEVSL HTEIKISSTD KAEITPATGA DTSDAFGNKK KQDRSSCGSN 540 TPSSSDVEAE HVPEKQDQVN EQEQQASCNN PSAGDTNHRR FRSIASTSES WKEVSEEGRM 600 AFDALFSRGK LPQSFSPPQA EGSKAVSKGE QSEATPVTVD LNKSASIIDH ELDTSVGPIA 660 SLPIELSHLN MKSRRTGFKP YKRCSVEAKE NRVPASDEVG TKRIRLDSEA ST* |
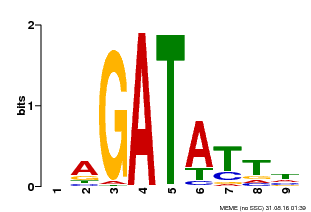
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Brast03G018100.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003573469.1 | 0.0 | protein CCA1 | ||||
| Refseq | XP_010234382.1 | 0.0 | protein CCA1 | ||||
| Refseq | XP_010234383.1 | 0.0 | protein CCA1 | ||||
| Refseq | XP_010234384.1 | 0.0 | protein CCA1 | ||||
| TrEMBL | I1I1F4 | 0.0 | I1I1F4_BRADI; Uncharacterized protein | ||||
| STRING | BRADI3G16515.1 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 3e-40 | circadian clock associated 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Brast03G018100.1.p |



