| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | AP2 | 59.6 | 7.7e-19 | 77 | 126 | 2 | 55 |
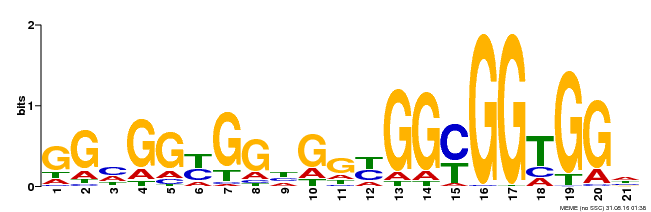
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
y+GVr++ +g+WvAeIr+p + r kr +lg+f ta eAa a+++a+++++g
Brast01G374200.1.p 77 AYRGVRQRT-WGKWVAEIREP---N-RgKRLWLGSFPTAVEAANAYDEAARAMYG 126
59****999.**********8...3.35************************998 PP
|
| Publications
? help Back to Top |
- Kikuchi S, et al.
Collection, mapping, and annotation of over 28,000 cDNA clones from japonica rice.
Science, 2003. 301(5631): p. 376-9
[PMID:12869764] - Sun SJ, et al.
Functional analysis of a novel Cys2/His2-type zinc finger protein involved in salt tolerance in rice.
J. Exp. Bot., 2010. 61(10): p. 2807-18
[PMID:20460361] - Mallikarjuna G,Mallikarjuna K,Reddy MK,Kaul T
Expression of OsDREB2A transcription factor confers enhanced dehydration and salt stress tolerance in rice (Oryza sativa L.).
Biotechnol. Lett., 2011. 33(8): p. 1689-97
[PMID:21528404] - Cui M, et al.
Induced over-expression of the transcription factor OsDREB2A improves drought tolerance in rice.
Plant Physiol. Biochem., 2011. 49(12): p. 1384-91
[PMID:22078375] - Yang A,Dai X,Zhang WH
A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice.
J. Exp. Bot., 2012. 63(7): p. 2541-56
[PMID:22301384] - Li X, et al.
LcSAIN1, a novel salt-induced gene from sheepgrass, confers salt stress tolerance in transgenic Arabidopsis and rice.
Plant Cell Physiol., 2013. 54(7): p. 1172-85
[PMID:23695503] - Schmidt R, et al.
Salt-responsive ERF1 regulates reactive oxygen species-dependent signaling during the initial response to salt stress in rice.
Plant Cell, 2013. 25(6): p. 2115-31
[PMID:23800963] - Huang YC,Huang WL,Hong CY,Lur HS,Chang MC
Comprehensive analysis of differentially expressed rice actin depolymerizing factor gene family and heterologous overexpression of OsADF3 confers Arabidopsis Thaliana drought tolerance.
Rice (N Y), 2012. 5(1): p. 33
[PMID:24279948] - Zhang XX, et al.
OsDREB2A, a rice transcription factor, significantly affects salt tolerance in transgenic soybean.
PLoS ONE, 2013. 8(12): p. e83011
[PMID:24376625] - Wang ST, et al.
MicroRNA319 positively regulates cold tolerance by targeting OsPCF6 and OsTCP21 in rice (Oryza sativa L.).
PLoS ONE, 2014. 9(3): p. e91357
[PMID:24667308] - Ma T, et al.
LcWRKY5: an unknown function gene from sheepgrass improves drought tolerance in transgenic Arabidopsis.
Plant Cell Rep., 2014. 33(9): p. 1507-18
[PMID:24913125] - Zhao J, et al.
Global transcriptional profiling of a cold-tolerant rice variety under moderate cold stress reveals different cold stress response mechanisms.
Physiol Plant, 2015. 154(3): p. 381-94
[PMID:25263631] - Chen M,Zhao Y,Zhuo C,Lu S,Guo Z
Overexpression of a NF-YC transcription factor from bermudagrass confers tolerance to drought and salinity in transgenic rice.
Plant Biotechnol. J., 2015. 13(4): p. 482-91
[PMID:25283804] - Gao Y, et al.
A maize phytochrome-interacting factor 3 improves drought and salt stress tolerance in rice.
Plant Mol. Biol., 2015. 87(4-5): p. 413-28
[PMID:25636202] - Cai W, et al.
Overexpression of Rat Neurons Nitric Oxide Synthase in Rice Enhances Drought and Salt Tolerance.
PLoS ONE, 2015. 10(6): p. e0131599
[PMID:26121399] - Sakuraba Y, et al.
Rice ONAC106 Inhibits Leaf Senescence and Increases Salt Tolerance and Tiller Angle.
Plant Cell Physiol., 2015. 56(12): p. 2325-39
[PMID:26443376] - Sato H, et al.
The Arabidopsis transcriptional regulator DPB3-1 enhances heat stress tolerance without growth retardation in rice.
Plant Biotechnol. J., 2016. 14(8): p. 1756-67
[PMID:26841113] - Yu Y, et al.
Transgenic rice expressing a cassava (Manihot esculenta Crantz) plasma membrane gene MePMP3-2 exhibits enhanced tolerance to salt and drought stresses.
Genet. Mol. Res., 2016.
[PMID:26909954]
|