 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009118853.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 334aa MW: 37140.8 Da PI: 8.8858 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 124.8 | 3.6e-39 | 188 | 265 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cq+e+C+adls+ak+yhrrhkvCe+hska++v+++gl+qrfCqqCsrfh lsefD++krsCr+rLa+hn+rrrk ++
XP_009118853.1 188 RCQAERCNADLSHAKHYHRRHKVCEFHSKASTVVAAGLSQRFCQQCSRFHLLSEFDNGKRSCRKRLADHNRRRRKCHQ 265
6*************************************************************************9865 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51141 | 31.77 | 186 | 263 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:4.10.1100.10 | 1.0E-33 | 186 | 250 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 9.32E-37 | 187 | 266 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 5.0E-29 | 189 | 262 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009554 | Biological Process | megasporogenesis | ||||
| GO:0009556 | Biological Process | microsporogenesis | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 334 aa Download sequence Send to blast |
MLDYEWDNTS AIMGSGEERH HHQDLDTTRA LSIFDPTQTN HHQSPYNNFS PPLFSNQQHL 60 TVVYGHQTAH NIPTNQFHHS FYDPNPYASS SSDSIYHAHS SASALFSFDQ SGPAGSGSSY 120 NFLVPKTEVD VVSRSLDFST SNRIGLNLGG RTYFSAGDDD FVSRLYRRSR PGELLGMGNS 180 LSTTLTPRCQ AERCNADLSH AKHYHRRHKV CEFHSKASTV VAAGLSQRFC QQCSRFHLLS 240 EFDNGKRSCR KRLADHNRRR RKCHQSASTS GKHIPDIITT PKSANDSGVK SSSASPSNAP 300 TPISLECFRQ RQFQTTPSSS TSAIWGSLAP AYNL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-31 | 178 | 262 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed during plant development. During pollen development, required for crosporogenesis that leads to archesporial formation and for the histogenesis of the microsporangia. During ovules development, involved in the transition to meiosis of the megaspore mother cells. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:12671094}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in shoot apical region and early floral tissues. Transcripts levels increase in developing pollen sacs, and decrease in later stage of anther development. Strongly expressed in the placental region of the carpels. {ECO:0000269|PubMed:12671094, ECO:0000269|PubMed:17093870}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. Binds specifically to the 5'-GTAC-3' core sequence. Involved in development and floral organogenesis. Required for ovule differentiation, pollen production, filament elongation, seed formation and siliques elongation. Also seems to play a role in the formation of trichomes on sepals. May positively modulate gibberellin (GA) signaling in flower. {ECO:0000269|PubMed:12671094, ECO:0000269|PubMed:16095614, ECO:0000269|PubMed:17093870}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00090 | SELEX | Transfer from AT1G02065 | Download |
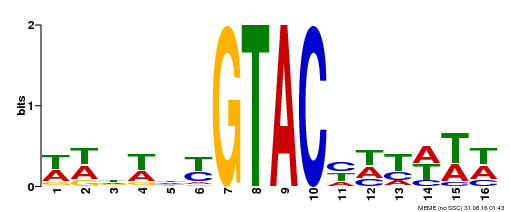
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009118853.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009118853.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 8 | ||||
| Swissprot | Q8GXL3 | 1e-120 | SPL8_ARATH; Squamosa promoter-binding-like protein 8 | ||||
| TrEMBL | A0A3P5YM77 | 0.0 | A0A3P5YM77_BRACM; Uncharacterized protein | ||||
| STRING | Bo8g118210.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM7010 | 27 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G02065.1 | 1e-113 | squamosa promoter binding protein-like 8 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



