 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009102986.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 390aa MW: 43666.6 Da PI: 8.4862 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 128.5 | 2.6e-40 | 171 | 247 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cq++gCe dls+ak+yhr+h+vCe hsk+p+v+vsg+e+rfCqqCsrfh++sefDe+krsCr+rL++hn+rrrk+q
XP_009102986.1 171 RCQIDGCELDLSSAKDYHRKHRVCENHSKCPKVTVSGIERRFCQQCSRFHAVSEFDEKKRSCRKRLSHHNARRRKPQ 247
6**************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 6.5E-33 | 164 | 233 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.317 | 169 | 246 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 3.14E-37 | 170 | 247 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 8.8E-31 | 172 | 245 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 390 aa Download sequence Send to blast |
MDCNMRSPFL WDLENLILPN SSKTENDKDQ LTTATEWETE KGEGIESMFP CFNGLERVSN 60 CSTTSLWHTP VSKSSQSTST NSSSPGVKQR MLASESSPGD SCCNIQVKAS VASAESDLSL 120 KLGKRTYSEE LWGRSNNNDI SAVSVKLLTP PSVVSRKKSK SSCGQSMQVP RCQIDGCELD 180 LSSAKDYHRK HRVCENHSKC PKVTVSGIER RFCQQCSRFH AVSEFDEKKR SCRKRLSHHN 240 ARRRKPQGVF PFNPDRVYDR RQHTNMLWNG LSLDTRSEEK FAWETTYDIK PTQTESGFTL 300 SFQRGNGSEQ QLFASSNHSY STYQTSGGLS AGKPKFQLHG QGVGEYSGVL HESQEFHRAL 360 SLLSTSSSDH LVQPHAQPLS LLFSYDGVPK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 4e-26 | 172 | 245 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 227 | 244 | KKRSCRKRLSHHNARRRK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bra.21592 | 0.0 | bud| flower | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed constitutively during plant development, weak increase during flowering. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:14573523}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00161 | DAP | Transfer from AT1G27360 | Download |
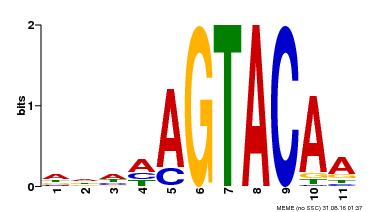
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009102986.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156 and miR157. {ECO:0000305|PubMed:12202040}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009102986.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 11 | ||||
| Refseq | XP_018508647.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 11 | ||||
| Swissprot | Q9FZK0 | 0.0 | SPL11_ARATH; Squamosa promoter-binding-like protein 11 | ||||
| TrEMBL | M4EMM0 | 0.0 | M4EMM0_BRARP; Uncharacterized protein | ||||
| STRING | Bra030040.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5004 | 16 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G27360.4 | 0.0 | squamosa promoter-like 11 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



