 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bathy05g02420 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Bathycoccus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 928aa MW: 101016 Da PI: 10.5492 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 49.9 | 7.5e-16 | 230 | 274 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+ E++++v+a k++G W++I +++g ++t+ q++s+ qk+
Bathy05g02420 230 RERWTEREHDRFVEALKLHGRA-WRKIEEHIG-TKTAVQIRSHAQKF 274
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.27E-15 | 224 | 280 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.178 | 225 | 279 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-7 | 227 | 276 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.8E-15 | 228 | 277 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 6.3E-12 | 229 | 277 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-12 | 230 | 273 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.77E-9 | 232 | 275 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010243 | Biological Process | response to organonitrogen compound | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043433 | Biological Process | negative regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:0043496 | Biological Process | regulation of protein homodimerization activity | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 928 aa Download sequence Send to blast |
MQRKVDKKKR FVWSFQIFRR DFLIQKVPLA FAPFNLRKNS RANDKRERRR QHLANARSRK 60 KEEQNVIYIP SDSNLFVMGD SSQQQQQYAL VAQMLAAQHR DQNVQNNNSA QRMMENNATA 120 QNNNANNTNT FHPQFPFVNY NLHGQPVVVN QMSQQQQQQQ YANNGKPTAN GQKSVLVGAT 180 MAPSEGHFLP QNTSASGYTL GGQQQPIPTH HTVNGQTVKV RKPYTITKQR ERWTEREHDR 240 FVEALKLHGR AWRKIEEHIG TKTAVQIRSH AQKFFAKLQK EHQKNLEADQ LSDVTRGGGT 300 TEDGATATRD TQTSGEGGLT QTTTENIKDA DTAKKISGNN NSKLLIQALM GKKSKSEVTG 360 NTKEEDEAEL LSNLAASGDK PATVKRTSSM STGGKTTASD IPPARPKRKP SHPYPRKQSS 420 LTEIQVRGVL PVGFVNVNNK KKGHSINNSL IDVANVEQHA SASAAQALMN ATSEMVRNRM 480 IQEMQMQHLL QQQQQQRQNN KNGNNNDTTA NPLLAHLAMQ QEVPNPLSQL HALMAANPMF 540 TAMNFGGGLP GMPNMAAVAA NSNGGSFIPS FATMNNNNNN NNAKLTTTTT TTDANNNKSG 600 TEEAKPTTTT TTDKTKNSEV SEDAKASLKP NSQSAFSGYV RPTTTTTTTN SNQYRQTGSA 660 ATTTTTANNI NNSSNNNNDS AVMAAVQQHQ QMQYWNAQMA ALAAMQDPNS FAQFLQHQQM 720 GNMAMAQQQR QQQNNIAYAA GVKSEAAAAT QEKNAAGEVK TARKAQAATQ TPAEATDAGK 780 TAKMIAASRK PNAPQVPKEI ELAVANKEKS PDSDKATEEE ENKNADNSNK DTITNTTTTT 840 TTTTTGRRSR RAAAVVAAEK VTKQVSGGRA TRRSNNGGSG SGSGSGSRNQ NQPGGNKNQG 900 SSGSDGSDDV AQKGFETTSF RVYQQEE* |
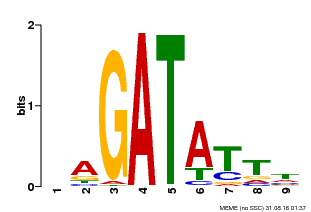
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FO082274 | 0.0 | FO082274.1 Bathycoccus prasinos genomic : chromosome_5. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007513081.1 | 0.0 | putative At5g37260-like protein | ||||
| TrEMBL | K8EF72 | 0.0 | K8EF72_9CHLO; Putative At5g37260-like protein | ||||
| STRING | XP_007513081.1 | 0.0 | (Bathycoccus prasinos) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP454 | 14 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 2e-31 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 19015745 |



