 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013610078.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | M-type_MADS | ||||||||
| Protein Properties | Length: 304aa MW: 34482 Da PI: 8.4786 | ||||||||
| Description | M-type_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 52.5 | 6.5e-17 | 10 | 51 | 2 | 43 |
---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TT CS
SRF-TF 2 rienksnrqvtfskRrngilKKAeELSvLCdaevaviifsst 43
+i n+s r+ tf kR++g++KK +ELS+LC+++++ ii+s+
XP_013610078.1 10 FIANDSSRKATFKKRKKGLIKKVNELSTLCGINACAIIYSPY 51
899*************************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 17.058 | 1 | 49 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 1.9E-24 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00266 | 4.63E-24 | 2 | 85 | No hit | No description |
| SuperFamily | SSF55455 | 4.45E-25 | 2 | 95 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.5E-8 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 1.2E-17 | 10 | 51 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.5E-8 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0043078 | Cellular Component | polar nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 304 aa Download sequence Send to blast |
MTRKKVKLAF IANDSSRKAT FKKRKKGLIK KVNELSTLCG INACAIIYSP YDTNPEVWPS 60 NSGVQRIISD FRKLPEMDQN KKMVDQEAFL RQRIAKASEN LKKQRKDNRE MEMTEVMFQC 120 LVGNMGMFNL NIMDLNDLGY MIDQYLKDVN RRIEILGSSG GGMELGETSN DAAAAPSEAA 180 GTLAIVPTTT DTIQPYHHQQ QHQMFRHHAA PHVGPYEQPL NLNLSHSQNQ QQGFTEMMMN 240 RPEQMSYAAE QMGFPFMNDN HHHHHYHHHQ QVPGESSTTP ATNVSASSAN PVTSSNPTNH 300 IWFR |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 2 | 24 | RKKVKLAFIANDSSRKATFKKRK |
| 2 | 21 | 25 | KKRKK |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the central cell of the female gametophyte and in early endosperm. Also detected in ovaries, young siliques, roots, leaves, stems, young flowers and anthers. {ECO:0000269|PubMed:16798889}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Controls central cell differentiation during female gametophyte development. Required for the expression of DEMETER and DD46, but not for the expression of FIS2 (PubMed:16798889). Probable transcription factor that may function in the maintenance of the proper function of the central cell in pollen tube attraction (Probable). {ECO:0000269|PubMed:16798889, ECO:0000305|PubMed:26462908}. | |||||
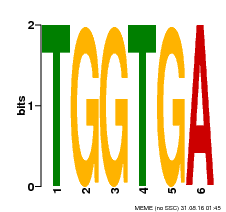
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00553 | DAP | Transfer from AT5G48670 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013610078.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009114607.1 | 0.0 | PREDICTED: agamous-like MADS-box protein AGL80 | ||||
| Refseq | XP_013610078.1 | 0.0 | PREDICTED: agamous-like MADS-box protein AGL80 | ||||
| Refseq | XP_013659196.1 | 0.0 | agamous-like MADS-box protein AGL80 | ||||
| Refseq | XP_013674953.1 | 0.0 | agamous-like MADS-box protein AGL80 | ||||
| Swissprot | Q9FJK3 | 1e-110 | AGL80_ARATH; Agamous-like MADS-box protein AGL80 | ||||
| TrEMBL | A0A078J738 | 0.0 | A0A078J738_BRANA; BnaAnng16690D protein | ||||
| TrEMBL | A0A0D3E8D3 | 0.0 | A0A0D3E8D3_BRAOL; Uncharacterized protein | ||||
| TrEMBL | A0A397XW25 | 0.0 | A0A397XW25_BRACM; Uncharacterized protein | ||||
| TrEMBL | A0A3P6E9T6 | 0.0 | A0A3P6E9T6_BRAOL; Uncharacterized protein | ||||
| TrEMBL | M4EL02 | 0.0 | M4EL02_BRARP; Uncharacterized protein | ||||
| STRING | Bra029469.1-P | 0.0 | (Brassica rapa) | ||||
| STRING | Bo9g078450.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM70 | 28 | 415 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G48670.1 | 1e-100 | AGAMOUS-like 80 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



