 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013604508.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1376aa MW: 153430 Da PI: 7.1694 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.2 | 0.0012 | 1259 | 1284 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C C++sFs+ +L H r+ +
XP_013604508.1 1259 YQCDmeGCTMSFSSEKQLSLHKRNiC 1284
99********************9877 PP
| |||||||
| 2 | zf-C2H2 | 16.4 | 2.5e-05 | 1284 | 1306 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk+F ++ +L++H+r+H
XP_013604508.1 1284 CPvkGCGKTFFSHKYLVQHQRVH 1306
9999*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 8.3E-17 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.243 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 4.1E-15 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 3.0E-53 | 202 | 371 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.783 | 205 | 371 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.25E-27 | 217 | 388 | No hit | No description |
| Pfam | PF02373 | 1.8E-37 | 235 | 354 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 0.001 | 1258 | 1281 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 9.4 | 1259 | 1281 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.339 | 1282 | 1311 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0015 | 1282 | 1306 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 7.2E-7 | 1283 | 1310 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1284 | 1306 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.3E-10 | 1298 | 1339 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 9.4E-10 | 1311 | 1335 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.801 | 1312 | 1341 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0015 | 1312 | 1336 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1314 | 1336 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.43E-8 | 1330 | 1364 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-9 | 1336 | 1365 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.388 | 1342 | 1373 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 2.2 | 1342 | 1368 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1344 | 1368 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1376 aa Download sequence Send to blast |
MAVSEQSQDV FPWLKSLPVA PEFKPTLAEF QDPMAYICKI EEEASRYGIC KIVPPVPPSS 60 KKTAINNLNR SLAARARARA RDGNGNGGKS DYDDGPTFTT RQQQIGFCPR RQRPVQRPVW 120 QSGERYTFDE FEFKAKNFEK SYLKKCGKKG SVSPLEVETL YWRASVDKPF SVEYANDMPG 180 SAFVPLSLAA ARRRESGGDC GTVGETAWNM RAMARAEGSL LKFMKEEIPG VTSPMVYIAM 240 MFSWFAWHVE DHDLHSLNYL HMGAAKTWYG VPKDAAMAFE EVVRVHGYGE ELNPLVTFST 300 LGEKTTVMSP EVFVRAGIPC CRLVQNPGDF VITFPGAYHS GFSHGFNFGE ASNIATPQWL 360 RMAKDAAIRR ASINYPPMVS HLQLLYDYAL ALGSRVPDSI HNKPRSSRLK DKKKSEGEKL 420 TNELFVQNVI HNNELLHSLG KGSPIALLPQ SSSDVSVCSD VRIGSHLGAN QGQTTLLIKS 480 EDLSSDSVMA SLSNGVKEKF TSLCERNRAD ETQGTSTDGE KRKNNGAVGL SDQRLFSCVT 540 CGVLSFDCVA IIQPKEAAAR YLMSADCSSL NDWTVASGSA NLGQDVVMPP SGKQDVGDLY 600 NAPVQTPYHS TTKTVDQRTS SSSLTKENGA LGLLASAYGD SSDSEEEDHK GLDNPVSDEV 660 EASSFVTDGN DEAGNGLSSA LNSQGLTCEK GKEVDVSHAN LSKGGNTSSV EITLPFIPRS 720 DDDFSRLHVF CLEHAAEVEQ RLHPIGGINI MLLCHPDYPR IEAEGKVVAK ELGVNHEWND 780 TEFKNVTRED EETIQAALAN VEAKAGNSDW AVKLGINLSH SAILSRSPLY SKQMPYNSVI 840 YNVFGRTSPA TPQVSGIRSS RQRKYVAGKW CGKVWMSHQV HPFLLEEDLE GEETERSHLR 900 AALDEDVTVH GNDSRDATTM FGRKYSRKRK ARGKAAPRKK LTSFKREAGV SDDTTSEDHS 960 YKQQWRAYGD EEESYFETGN EVSGDSSNQM SDQQQLKVVE SDDDEVSERS LGQEYAVRRE 1020 YASSESSMEN GFQVYREDQP MYGDDDMYRH PRGIPRNKRT KVFRDLVSYD SEDQQRERVF 1080 TSDAQTSRMG DEYDSEENSL EEQDFCSSGK RQTRSIAKRK VKTKIVQSLG DTGGRTLLQS 1140 GSRKKMKELD SYMEGPSTRL RVRTPKPSRG SSATKPKKTG KKGRNVSFSE VASEEEVEEN 1200 EEESEEASTR IRVRTLKPPR RSLETKPKKT GKKGSNVSLS GVASEEEVEE NEQEECLPYQ 1260 CDMEGCTMSF SSEKQLSLHK RNICPVKGCG KTFFSHKYLV QHQRVHSDDR PLKCPWKGCK 1320 MTFKWAWSRT EHIRVHTGER PYICAEPGCS QTFRFVSDFS RHKRKTGHSA KKIKKK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 1e-73 | 1256 | 1376 | 20 | 140 | Lysine-specific demethylase REF6 |
| 6a58_A | 1e-73 | 1256 | 1376 | 20 | 140 | Lysine-specific demethylase REF6 |
| 6a59_A | 1e-73 | 1256 | 1376 | 20 | 140 | Lysine-specific demethylase REF6 |
| 6ip0_A | 6e-70 | 9 | 394 | 4 | 355 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 6e-70 | 9 | 394 | 4 | 355 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 927 | 939 | KRKARGKAAPRKK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bol.10792 | 0.0 | leaf | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in the shoot apical meristem and primary and secondary root tips, and lower expression in cotyledons, leaves and root axis along vascular tissues. Detected in inflorescences, stems and siliques. {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18713399}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
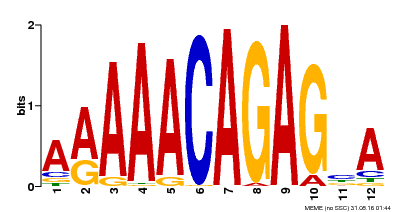
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013604508.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013604508.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 isoform X2 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A0D3DRD2 | 0.0 | A0A0D3DRD2_BRAOL; Uncharacterized protein | ||||
| STRING | Bo8g076910.1 | 0.0 | (Brassica oleracea) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



