 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013591866.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 633aa MW: 70282.6 Da PI: 6.1454 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 97.2 | 1.5e-30 | 56 | 140 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+++e+laL+++r+em++++r++ lk+plWe+vs+k+ e g++rs+k+Ckek+en++k+yk++ke++++r++++ +++f+qlea
XP_013591866.1 56 RWPREETLALLRIRSEMDSTFRDATLKAPLWEHVSRKLLELGYKRSAKKCKEKFENVQKYYKRTKETRGGRHDGK--AYKFFSQLEA 140
8*********************************************************************86555..5******985 PP
| |||||||
| 2 | trihelix | 104.1 | 1e-32 | 406 | 490 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
rW+k e+laLi++r+ me r++++ k+ lWee+s+ m++ g++r++k+Ckekwen+nk+ykk+ke++k+r +++ +tcpyf++l+
XP_013591866.1 406 RWPKAEILALINLRSGMEPRYQDNVPKGLLWEEISSSMKRMGYNRNAKRCKEKWENINKYYKKVKESNKER-PQDAKTCPYFHRLD 490
8*********************************************************************8.99999*******97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50090 | 7.027 | 49 | 113 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.0031 | 53 | 115 | IPR001005 | SANT/Myb domain |
| CDD | cd12203 | 1.37E-27 | 55 | 120 | No hit | No description |
| Pfam | PF13837 | 9.4E-21 | 55 | 141 | No hit | No description |
| PROSITE profile | PS50090 | 6.922 | 399 | 463 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.0045 | 403 | 465 | IPR001005 | SANT/Myb domain |
| Pfam | PF13837 | 7.4E-22 | 405 | 490 | No hit | No description |
| CDD | cd12203 | 2.61E-30 | 406 | 470 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:0010090 | Biological Process | trichome morphogenesis | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0032876 | Biological Process | negative regulation of DNA endoreduplication | ||||
| GO:0042631 | Biological Process | cellular response to water deprivation | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:2000037 | Biological Process | regulation of stomatal complex patterning | ||||
| GO:2000038 | Biological Process | regulation of stomatal complex development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 633 aa Download sequence Send to blast |
MEQGAGGGNE VVEEASPISS RPPANMEELM RFSAAADEGG GLGGGGSSAS SSSGNRWPRE 60 ETLALLRIRS EMDSTFRDAT LKAPLWEHVS RKLLELGYKR SAKKCKEKFE NVQKYYKRTK 120 ETRGGRHDGK AYKFFSQLEA LNTTPPSSSL DATPLSVANP IQPPPSSSQF PVFPLPQTLP 180 HSVSFTPNVA PPPPPPPAPM GPTFPGVTFS SHSSSTASGM GSDDDDDEED IMDVDQAGPS 240 SRKRKRGNRG GGKMMELFDG LVRQVMQKQA SMQRSFLEAL EKREQERLHR EEAWKRQEMS 300 RLAREHEIMS QERAASASRD AAIISMIQKI TGHTIQLPPS FSSQPSPPPR QPPPAAKRPS 360 SQPVEPPQLQ PIMAIPQQQV LPPPPQPPPQ QEVTMSSDQS SPSSSRWPKA EILALINLRS 420 GMEPRYQDNV PKGLLWEEIS SSMKRMGYNR NAKRCKEKWE NINKYYKKVK ESNKERPQDA 480 KTCPYFHRLD LLYRNKVLGS GGSSSASALP HQDQISTVQK QSPVSAVKPP QGVVTVGSAS 540 SEGEEPREES PQETEKPEDL VMKELMQHQQ QDSMISEYEK IEESHNYNNM EEEEMDEELD 600 EDEKSAAYEV AFQSPANRGG NGHTEPPFLT MVQ |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 98 | 106 | KRSAKKCKE |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Mostly expressed in siliques, and, to a lower extent, in growing root hairs, leaves, stems, and flowers. Present in abaxial epidermal cells, predominantly in guard cells, pavement cells, and meristemoids. {ECO:0000269|PubMed:21169508, ECO:0000269|PubMed:9501260}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription repressor that binds specific DNA sequence such as GT3 box 5'-GGTAAA-3' in the SDD1 promoter. Negative regulator of water use efficiency (WUE) via the promotion of stomatal density and distribution by the transcription repression of SDD1. Regulates the expression of several cell cycle genes and endoreduplication, especially in trichomes where it prevents ploidy-dependent plant cell growth. {ECO:0000269|PubMed:19717615, ECO:0000269|PubMed:21169508}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00177 | DAP | Transfer from AT1G33240 | Download |
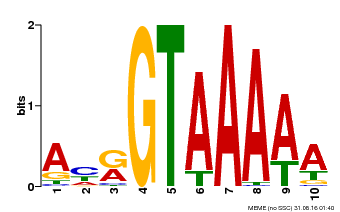
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013591866.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by water stress. {ECO:0000269|PubMed:21169508}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC021045 | 1e-120 | AC021045.2 Arabidopsis thaliana chromosome I BAC T9L6 genomic sequence, complete sequence. | |||
| GenBank | AC027035 | 1e-120 | AC027035.5 Arabidopsis thaliana chromosome 1 BAC T16O9 genomic sequence, complete sequence. | |||
| GenBank | AJ003215 | 1e-120 | AJ003215.1 Arabidopsis thaliana GTL1 gene. | |||
| GenBank | AY140085 | 1e-120 | AY140085.1 Arabidopsis thaliana DNA-binding factor, putative (At1g33240) mRNA, complete cds. | |||
| GenBank | BT008824 | 1e-120 | BT008824.1 Arabidopsis thaliana At1g33240 mRNA, complete cds. | |||
| GenBank | CP002684 | 1e-120 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013591866.1 | 0.0 | PREDICTED: trihelix transcription factor GTL1 | ||||
| Swissprot | Q9C882 | 0.0 | GTL1_ARATH; Trihelix transcription factor GTL1 | ||||
| TrEMBL | A0A0D3CS58 | 0.0 | A0A0D3CS58_BRAOL; Uncharacterized protein | ||||
| STRING | Bo6g049430.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6226 | 27 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G33240.1 | 4e-90 | GT-2-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



