 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSBRNA2T00110242001 | ||||||||
| Common Name | GSBRNA2T00110242001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 674aa MW: 74578.6 Da PI: 7.0539 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.3 | 6.3e-19 | 42 | 95 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k +++t++q++eLe++F+ +++p++++r eL +kl+L+ +q+k+WFqNrR+++k
GSBRNA2T00110242001 42 KYRRHTSYQIQELESFFKVCPHPNEKQRLELGSKLSLESKQIKFWFQNRRTQMK 95
56789**********************************************999 PP
| |||||||
| 2 | START | 170.2 | 1.3e-53 | 207 | 416 | 3 | 205 |
HHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECTT CS
START 3 aeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetla....kaetlevissg 88
a a++el+++a+++ p+W+k s + + ++++ + + +r++g+v ++ +lve+l+d++ +W e+++ a+t+evis+g
GSBRNA2T00110242001 207 AAVAMDELIRLAEVDNPLWTKCS--KSERDSMKHDQYTSI-FAGSSRETGLVLINSSTLVETLMDTN-RWAEMFEsivaVASTVEVISNG 292
56689******************..666666666655444.5567**********************.**********9*********** PP
......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE CS
START 89 ......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvd 171
gal lm+ae+q++splvp R + f+Ry++q+g+g w++vdvS d++++ + +s+ ++lpSg++i++ +ng skvtw+eh +
GSBRNA2T00110242001 293 sggsrnGALLLMQAEFQVMSPLVPiRQVKFLRYCKQHGDGLWAVVDVSYDVNRESQDLKSYGGLKRLPSGCIIQDIGNGCSKVTWIEHSE 382
******************************************************99999******************************* PP
--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 172 lkgrlphwllrslvksglaegaktwvatlqrqce 205
++g+++h l+++l++s++ ga +w+atlqr ce
GSBRNA2T00110242001 383 YEGSHIHPLYQQLLGSSVGLGATKWLATLQRRCE 416
*********************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.84E-18 | 22 | 97 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-19 | 25 | 97 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.406 | 37 | 97 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.7E-15 | 39 | 101 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.62E-15 | 42 | 97 | No hit | No description |
| Pfam | PF00046 | 1.2E-16 | 42 | 95 | IPR001356 | Homeobox domain |
| PROSITE profile | PS50848 | 43.912 | 196 | 420 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.43E-31 | 200 | 418 | No hit | No description |
| CDD | cd08875 | 1.87E-100 | 201 | 416 | No hit | No description |
| SMART | SM00234 | 4.0E-40 | 205 | 417 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 1.1E-4 | 205 | 400 | IPR023393 | START-like domain |
| Pfam | PF01852 | 2.1E-47 | 207 | 417 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 7.58E-19 | 443 | 666 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 674 aa Download sequence Send to blast |
MNGDFDISKG DDEFESDSFE AMSGDDDDKH EQRPKKKKKM SKYRRHTSYQ IQELESFFKV 60 CPHPNEKQRL ELGSKLSLES KQIKFWFQNR RTQMKTQLER HENAILRQEN EKLRVENGIL 120 KEAMRSPPTC NNCGGAATPG EVSHEQQQLR MENAKLKYEL DKLCALSNRF IGGSISLEQP 180 SNGGVGSQDL SLGHGFTRGT STFMDIAAVA MDELIRLAEV DNPLWTKCSK SERDSMKHDQ 240 YTSIFAGSSR ETGLVLINSS TLVETLMDTN RWAEMFESIV AVASTVEVIS NGSGGSRNGA 300 LLLMQAEFQV MSPLVPIRQV KFLRYCKQHG DGLWAVVDVS YDVNRESQDL KSYGGLKRLP 360 SGCIIQDIGN GCSKVTWIEH SEYEGSHIHP LYQQLLGSSV GLGATKWLAT LQRRCESYTT 420 LLSSQDQTGL PLAGTKSTLT LAQRMKRNFY SGITGSPIHK WEKLVAENVG QETRILTRKS 480 LQPSGVVLSA ATSMWLPVTQ QRLFEFLCDS KCRNQWDILC NGASMENMLL IPQRQSEGRC 540 VSLLQHAGKH QNESSMLILQ ETWSDASGAL VVYAPVDVPS MNMVMSGGDS ANVALLPSGF 600 SISPDGSSSW SDQIDKNGGL VNHESKGCLL TVGFQILVNS VPTTKLNMES VQTVNNLIAC 660 TIHKIKAALS IPA* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in cells around the base of leaf primordia, in the outermost 2 to 3 cell layers along the boundary between two leaf primordia. Expressed in lateral root primordia and tips, and in the epidermal boundaries of two cotyledons at heart-stage embryo. {ECO:0000269|PubMed:16778018}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds to the DNA sequence 5'-GCATTAAATGC-3'. {ECO:0000269|PubMed:16778018}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00559 | DAP | Transfer from AT5G52170 | Download |
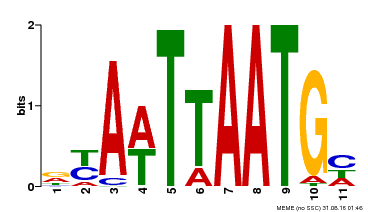
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GSBRNA2T00110242001 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB025603 | 1e-105 | AB025603.1 Arabidopsis thaliana genomic DNA, chromosome 5, BAC clone:F17P19. | |||
| GenBank | CP002688 | 1e-105 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013686466.1 | 0.0 | homeobox-leucine zipper protein HDG7-like | ||||
| Swissprot | Q9LTK3 | 0.0 | HDG7_ARATH; Homeobox-leucine zipper protein HDG7 | ||||
| TrEMBL | A0A3P6CY22 | 0.0 | A0A3P6CY22_BRACM; Uncharacterized protein | ||||
| STRING | Bra028295.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52170.1 | 0.0 | homeodomain GLABROUS 7 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



