 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G52170.1 | ||||||||
| Common Name | F17P19.7, HDG7, HDGL2-7 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 682aa MW: 75638.1 Da PI: 6.0005 | ||||||||
| Description | homeodomain GLABROUS 7 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.2 | 6.6e-19 | 59 | 113 | 2 | 56 |
T--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 2 rkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k +++t++q++eLe++F+++++p++++r eL kkl L+ +q+k+WFqNrR+++k
AT5G52170.1 59 TKYHRHTSYQIQELESFFKECPHPNEKQRLELGKKLTLESKQIKFWFQNRRTQMK 113
678899**********************************************999 PP
| |||||||
| 2 | START | 154.7 | 7.7e-49 | 224 | 426 | 2 | 206 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECTT......E CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetla....kaetlevissg......g 89
la ea++el+k+a++e +W++ s ++ ++ f +r++g+v ++ lve+l+d++ +W e+++ a+tlevis+g g
AT5G52170.1 224 LAMEAMDELLKLAELETSLWSSKS----EKGSMNHF--------PGSRETGLVLINSLALVETLMDTN-KWAEMFEcivaVASTLEVISNGsdgsrnG 308
57799***************9999....77788888........459*********************.***************************** PP
EEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHH CS
START 90 alqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvk 186
+ lm+ae+q++splvp + f+Ry++q+g+g w++vdvS d ++ +++ +s+ ++++pSg++i++ +ng skvtw+eh ++++++ h+l+++l++
AT5G52170.1 309 SILLMQAEFQVMSPLVPiKQKKFLRYCKQHGDGLWAVVDVSYDINRGNENLKSYGGSKMFPSGCIIQDIGNGCSKVTWIEHSEYEESHTHSLYQPLLS 406
************************************************************************************************** PP
HHHHHHHHHHHHHTXXXXXX CS
START 187 sglaegaktwvatlqrqcek 206
s++ ga +w+atlqrqce+
AT5G52170.1 407 SSVGLGATKWLATLQRQCES 426
******************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.25E-18 | 38 | 115 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 6.8E-20 | 43 | 115 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 5.2E-14 | 53 | 119 | IPR001356 | Homeobox domain |
| PROSITE profile | PS50071 | 16.973 | 55 | 115 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.1E-16 | 59 | 113 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.29E-15 | 62 | 115 | No hit | No description |
| PROSITE profile | PS50848 | 41.437 | 214 | 429 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.51E-30 | 220 | 427 | No hit | No description |
| CDD | cd08875 | 2.81E-103 | 220 | 425 | No hit | No description |
| SMART | SM00234 | 8.8E-37 | 223 | 426 | IPR002913 | START domain |
| Pfam | PF01852 | 1.9E-42 | 224 | 426 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.47E-17 | 450 | 674 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000016 | anatomy | lateral root primordium | ||||
| PO:0000017 | anatomy | vascular leaf primordium | ||||
| PO:0000026 | anatomy | primary root tip | ||||
| PO:0005679 | anatomy | epidermis | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0025257 | anatomy | primary root elongation zone | ||||
| PO:0007131 | developmental stage | seedling development stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 682 aa Download sequence Send to blast |
MNGDLEVDMS RGDFNPSFFL GKLKDDEFES RSLSDDSFDA MSGDEDKQEQ RPKKKKRKTK 60 YHRHTSYQIQ ELESFFKECP HPNEKQRLEL GKKLTLESKQ IKFWFQNRRT QMKTQLERHE 120 NVILKQENEK LRLENSFLKE SMRGSLCIDC GGAVIPGEVS FEQHQLRIEN AKLKEELDRI 180 CALANRFIGG SISLEQPSNG GIGSQHLPIG HCVSGGTSLM FMDLAMEAMD ELLKLAELET 240 SLWSSKSEKG SMNHFPGSRE TGLVLINSLA LVETLMDTNK WAEMFECIVA VASTLEVISN 300 GSDGSRNGSI LLMQAEFQVM SPLVPIKQKK FLRYCKQHGD GLWAVVDVSY DINRGNENLK 360 SYGGSKMFPS GCIIQDIGNG CSKVTWIEHS EYEESHTHSL YQPLLSSSVG LGATKWLATL 420 QRQCESFTML LSSEDHTGLS HAGTKSILKL AQRMKLNFYS GITASCIHKW EKLLAENVGQ 480 DTRILTRKSL EPSGIVLSAA TSLWLPVTQQ RLFEFLCDGK CRNQWDILSN GASMENTLLV 540 PKGQQEGSCV SLLRAAGNDQ NESSMLILQE TWNDVSGALV VYAPVDIPSM NTVMSGGDSA 600 YVALLPSGFS ILPDGSSSSS DQFDTDGGLV NQESKGCLLT VGFQILVNSL PTAKLNVESV 660 ETVNNLIACT IHKIRAALRI PA |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 49 | 57 | QRPKKKKRK |
| 2 | 52 | 57 | KKKKRK |
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 18423360 | 0.0 | ||||
| Genevisible | 248370_at | 0.0 | ||||
| Expression Atlas | AT5G52170 | - | ||||
| AtGenExpress | AT5G52170 | - | ||||
| ATTED-II | AT5G52170 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in cells around the base of leaf primordia, in the outermost 2 to 3 cell layers along the boundary between two leaf primordia. Expressed in lateral root primordia and tips, and in the epidermal boundaries of two cotyledons at heart-stage embryo. {ECO:0000269|PubMed:16778018}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a homeobox-leucine zipper family protein belonging to the HD-ZIP IV family. | |||||
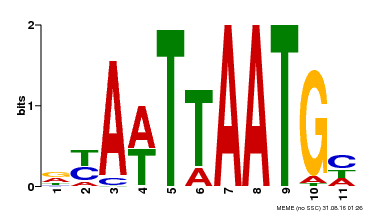
| UniProt | Probable transcription factor that binds to the DNA sequence 5'-GCATTAAATGC-3'. {ECO:0000269|PubMed:16778018}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00559 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G52170.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G52170 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB025603 | 0.0 | AB025603.1 Arabidopsis thaliana genomic DNA, chromosome 5, BAC clone:F17P19. | |||
| GenBank | CP002688 | 0.0 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001318786.1 | 0.0 | homeodomain GLABROUS 7 | ||||
| Refseq | NP_200030.1 | 0.0 | homeodomain GLABROUS 7 | ||||
| Swissprot | Q9LTK3 | 0.0 | HDG7_ARATH; Homeobox-leucine zipper protein HDG7 | ||||
| TrEMBL | A0A1P8BG00 | 0.0 | A0A1P8BG00_ARATH; Homeodomain GLABROUS 7 | ||||
| STRING | AT5G52170.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM491 | 28 | 149 | Representative plant | OGRP145 | 15 | 136 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G52170.1 |
| Entrez Gene | 835293 |
| iHOP | AT5G52170 |
| wikigenes | AT5G52170 |



