 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bradi3g34200.1.p | ||||||||
| Common Name | BRADI_3g34200, LOC100836477 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 707aa MW: 76240.2 Da PI: 6.7157 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39 | 1.4e-12 | 529 | 574 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr ++P+ K++Ka+ L A+ YI++L
Bradi3g34200.1.p 529 NHVEAERQRREKLNQRFYALRAVVPNV-----SKMDKASLLGDAISYINEL 574
799***********************6.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.1E-51 | 52 | 234 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.39 | 525 | 574 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 7.85E-18 | 528 | 596 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.71E-14 | 528 | 575 | No hit | No description |
| Pfam | PF00010 | 4.7E-10 | 529 | 574 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.5E-18 | 529 | 596 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 9.5E-15 | 531 | 580 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009269 | Biological Process | response to desiccation | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009867 | Biological Process | jasmonic acid mediated signaling pathway | ||||
| GO:0009963 | Biological Process | positive regulation of flavonoid biosynthetic process | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0043619 | Biological Process | regulation of transcription from RNA polymerase II promoter in response to oxidative stress | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051090 | Biological Process | regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:2000068 | Biological Process | regulation of defense response to insect | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 707 aa Download sequence Send to blast |
MNLWTDDNAS MMEAFMASAA DLPTFPWGAA AATPPPPAAV MPQQPAFNQD TLQQRLQAII 60 EGSRETWTYA IFWQSSTDAG AGASLLGWGD GYYKGCDDAD KRARQQPTPA SAAEQEHRKR 120 VLRELNSLIA GGGAAAPDEA VEEEVTDTEW FFLVSMTQSF PNGMGLPGQA LYTRQPTWIA 180 SGLASAPCER ARQAYTFGLR TMVCIPVGTG VLELGATEVI FQTADSLGRI RSLFNLNGGG 240 GGGGAGSSWP PVAPHQQHGG DQAETDPSVL WLTDAPVGDM KESPSVEISV SKPPPPPQIH 300 HFENGSTSTL TENAGPSLHA HQQPATLAPA APPRQNQHPH QLQLQHQQSQ QQQQQQQGPF 360 RRELNFSDFA TNASVTVTPP FFKPESGEIL NFGADSTSRR NPSPAPPAAA ASLTTAPGSL 420 FSQHTATVTA PTNEAKNNPK RSMEATSRAS NTNHHPSATA NEGMLSFSSA PTTRPSTGTG 480 APAKSESDHS DLEASVREVE SSRVVPPPEE KRPRKRGRKP ANGREEPLNH VEAERQRREK 540 LNQRFYALRA VVPNVSKMDK ASLLGDAISY INELRGKMTA LESDKDTLHS QIEALKKERD 600 ARPVAPLSGV HDSGPRCHAV EIEAKILGLE AMIRVQCHKR NHPAAKLMTA LRELDLDVYH 660 ASVSVVKDIM IQQVAVKMPN RVYSQDQLNA ALYSRLAEPG APVPIR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rqw_A | 4e-65 | 44 | 237 | 1 | 195 | Transcription factor MYC3 |
| 4rqw_B | 4e-65 | 44 | 237 | 1 | 195 | Transcription factor MYC3 |
| 4rs9_A | 4e-65 | 44 | 237 | 1 | 195 | Transcription factor MYC3 |
| 4yz6_A | 4e-65 | 44 | 237 | 1 | 195 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 510 | 518 | KRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in jasmonate (JA) signaling pathway during spikelet development. Binds to the G2 region G-box (5'-CACGTG-3') of the MADS1 promoter and thus directly regulates the expression of MADS1. Its function in MADS1 activation is abolished by TIFY3/JAZ1 which directly target MYC2 during spikelet development. {ECO:0000269|PubMed:24647160}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00084 | PBM | Transfer from AT1G32640 | Download |
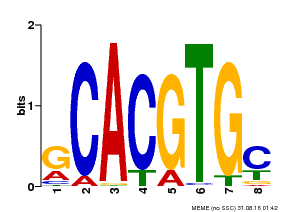
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bradi3g34200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK357816 | 0.0 | AK357816.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1062G16. | |||
| GenBank | AK359813 | 0.0 | AK359813.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1103K10. | |||
| GenBank | AK360465 | 0.0 | AK360465.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1118K08. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003574381.1 | 0.0 | transcription factor MYC2 | ||||
| Swissprot | Q336P5 | 0.0 | MYC2_ORYSJ; Transcription factor MYC2 | ||||
| TrEMBL | I1I6F3 | 0.0 | I1I6F3_BRADI; Uncharacterized protein | ||||
| STRING | BRADI3G34200.1 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4452 | 31 | 57 | Representative plant | OGRP3733 | 12 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G32640.1 | 1e-140 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bradi3g34200.1.p |
| Entrez Gene | 100836477 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



