 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm_27.model.AmTr_v1.0_scaffold00038.113 | ||||||||
| Common Name | AMTR_s00038p00173360 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; basal Magnoliophyta; Amborellales; Amborellaceae; Amborella
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1137aa MW: 128031 Da PI: 5.6892 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 179.5 | 4e-56 | 46 | 161 | 3 | 118 |
CG-1 3 kekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhe 71
+ ++rwl++ e+++iL n++++++++++++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktvrE+he
evm_27.model.AmTr_v1.0_scaffold00038.113 46 EAQNRWLRPAEVCEILRNYHNFHIASDPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVREAHE 114
5599***************************************************************** PP
CG-1 72 kLKvggvevlycyYahseenptfqrrcywlLeeelekivlvhylevk 118
+LK g ++vl+cyYah+een++fqrr+ywlLeeele+ivlvhy+evk
evm_27.model.AmTr_v1.0_scaffold00038.113 115 RLKAGRIDVLHCYYAHGEENENFQRRSYWLLEEELEHIVLVHYREVK 161
********************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.074 | 40 | 166 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.0E-75 | 43 | 161 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.6E-50 | 46 | 159 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 8.9E-6 | 553 | 638 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.1E-6 | 553 | 627 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 2.8E-17 | 553 | 638 | IPR014756 | Immunoglobulin E-set |
| CDD | cd00204 | 6.08E-13 | 740 | 849 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 7.9E-16 | 740 | 852 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 16.016 | 746 | 851 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.38E-15 | 748 | 851 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.16 | 790 | 819 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 500 | 829 | 858 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.84 | 965 | 987 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 4.78E-7 | 966 | 1016 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 8.206 | 968 | 993 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0015 | 968 | 986 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0096 | 988 | 1010 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.176 | 989 | 1013 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0012 | 991 | 1010 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1137 aa Download sequence Send to blast |
MLACFNLFAS QINCIRSVGA YGCAVMAESR HYALSNPLDI SQIVLEAQNR WLRPAEVCEI 60 LRNYHNFHIA SDPPNRPPSG SLFLFDRKVL RYFRKDGHNW RKKKDGKTVR EAHERLKAGR 120 IDVLHCYYAH GEENENFQRR SYWLLEEELE HIVLVHYREV KGNKTGYGRS RDAEKTFQVT 180 PTSSPVHSAS LNSNPSQLHS QTTPGSSMSI GQSEYEDAES GNPQVTSRYK SLLELQQPEY 240 RLQRNQKDAD LLNSYLEVLR TDNIFKSHPL IFTSKGNHHD NQSAAPEMSF VSHDRNNVLE 300 EKNIGGFEMN QLEPRKQMDM ASWSDVLGHG TMGSSDKSVY VGGLPNKQFN GIFEQLFAED 360 ISTKSEALAK PYAQEEWQIA SSEDSSKATA NTRIHTEQGS EPCGNYQQSK YLWMKPHIDQ 420 EPFSIQFGNL KDSCIILKDG SFPEVGHFQE SKSNEDEVGV EEYAVHSRFP EQPLLKSLSK 480 TEGEEGLKKL DSFSRWMSNE FGGEDVVVSS ESRSFWSTLD STDVVDDSRM PHQLNLGTDS 540 LSPSISQDQL FSIIDFSPTW AYSGLDCKVL ITGTFLMNQN QVEKCQWSCM FGEVEVPAQV 600 LTENVLRCHT PSHASGRVPF YVTCSNRVAC SEIREFEFLD CAPEYMDTFT DIDNTSTNEM 660 VLRVRLASLL SLGSSIPVKS LSSNVREETY ISGKINSLLK DNDDEWFQIE NLTDDEDLFP 720 GKAKDQLVQK LLKEKLHAWL LVKAGEDGKG PNVLDTQGQG VLHLTSALGY DWAIAPIVAA 780 GVNINFRDVS GWTALHWAAS CGRERTVAAI IALGGAPGAL SDPTPKFSSG QTPADLASVN 840 GHKGIAGYLA ESALTSHLSK LTIEEAIEDG NELALTSENA LEPTNDEIID QFNDGDSLDG 900 LSLRNSLTAV RNAAQAAARI HEVFRVQSFH RKKLIEYGDD KFGMSDERAL SLISVQKMRK 960 TGNDEPVHAA VRIQRKFRGW KGRKEFLVIR QRIVHLQAFF RGYQVRKHYK KIIWSVGIVE 1020 KAILRWRRKG SGLRGFKPEA SIEGPNAQAE SSQSDDYDFL KVGRRQTEER LDKALARVQS 1080 MVQYPEARAQ YRRLMNVVNE FQESKVDSER LLRQAEEIEY ELIDCVTLEE EDTIMG* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
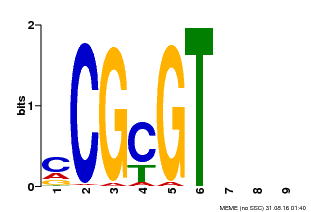
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006853146.3 | 0.0 | calmodulin-binding transcription activator 2 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | U5D2P6 | 0.0 | U5D2P6_AMBTC; Uncharacterized protein | ||||
| STRING | ERN14613 | 0.0 | (Amborella trichopoda) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP562 | 15 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm_27.model.AmTr_v1.0_scaffold00038.113 |



