 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT3G02940.1 | ||||||||
| Common Name | AtMYB107, F13E7.11, MYB107 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 321aa MW: 36627.9 Da PI: 6.6732 | ||||||||
| Description | myb domain protein 107 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 58.5 | 1.5e-18 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd++l++ ++++G g+W++ ++ g++R++k+c++rw +yl
AT3G02940.1 14 KGPWTPEEDQKLINHIRKHGHGSWRALPKQAGLNRCGKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 57.4 | 3.5e-18 | 67 | 111 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rg++T+eE++ +++++ +lG++ W++Ia +++ gRt++++k++w+++
AT3G02940.1 67 RGNFTAEEEQTIINLHSLLGNK-WSSIAGHLP-GRTDNEIKNYWNTH 111
89********************.*********.************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-27 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.763 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.42E-32 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.0E-16 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.7E-17 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.46E-12 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 27.394 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.7E-28 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.2E-16 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.0E-16 | 67 | 111 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.85E-12 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006357 | Biological Process | regulation of transcription from RNA polymerase II promoter | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000981 | Molecular Function | RNA polymerase II transcription factor activity, sequence-specific DNA binding | ||||
| GO:0001135 | Molecular Function | transcription factor activity, RNA polymerase II transcription factor recruiting | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 321 aa Download sequence Send to blast |
MGRSPCCDES GLKKGPWTPE EDQKLINHIR KHGHGSWRAL PKQAGLNRCG KSCRLRWTNY 60 LRPDIKRGNF TAEEEQTIIN LHSLLGNKWS SIAGHLPGRT DNEIKNYWNT HIRKKLIQMG 120 IDPVTHRPRT DHLNVLAALP QLLAAANFNN LLNLNQNIQL DATSVAKAQL LHSMIQVLSN 180 NNTSSSFDIH HTTNNLFGQS SFLENLPNIE NPYDQTQGLS HIDDQPLDSF SSPIRVVAYQ 240 HDQNFIPPLI STSPDESKET QMMVKNKEIM KYNDHTSNPS STSTFTQDHQ PWCDIIDDEA 300 SDSYWKEIIE QTCSEPWPFR E |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 7e-30 | 12 | 116 | 5 | 108 | B-MYB |
| 1h8a_C | 1e-29 | 12 | 116 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.14734 | 0.0 | root| silique | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 258620_at | 0.0 | ||||
| Expression Atlas | AT3G02940 | - | ||||
| AtGenExpress | AT3G02940 | - | ||||
| ATTED-II | AT3G02940 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Specifically and transiently expressed in root endodermal cells overlying the early stages of lateral root primordia formation. {ECO:0000269|PubMed:24902892, ECO:0000269|PubMed:25482809}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a putative transcription factor (MYB107). | |||||
| UniProt | Transcription factor that acts as negative regulator of lateral root (LR) development. Required for normal auxin responses during LR development. May be part of a negative feedback loop stimulated specifically in the endodermis upon LR initiation to ensure that LRs are formed only in the correct place. {ECO:0000269|PubMed:24902892}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00327 | DAP | 27203113 | Download |
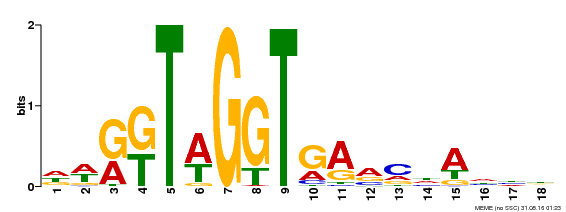
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT3G02940.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by abscisic acid (ABA), auxin and gravity in roots. {ECO:0000269|PubMed:24902892}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- Hormone ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hormone | |||||
| AHD | salicylic acid | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT3G02940 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF249310 | 0.0 | AF249310.2 Arabidopsis thaliana putative transcription factor (MYB107) mRNA, complete cds. | |||
| GenBank | AY519583 | 0.0 | AY519583.1 Arabidopsis thaliana MYB transcription factor (At3g02940) mRNA, complete cds. | |||
| GenBank | BT025660 | 0.0 | BT025660.1 Arabidopsis thaliana At3g02940 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001326399.1 | 0.0 | myb domain protein 107 | ||||
| Refseq | NP_186944.1 | 0.0 | myb domain protein 107 | ||||
| Swissprot | Q9S9Z2 | 2e-90 | MYB93_ARATH; Transcription factor MYB93 | ||||
| TrEMBL | Q9LDI5 | 0.0 | Q9LDI5_ARATH; At3g02940 | ||||
| STRING | AT3G02940.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 | Representative plant | OGRP5 | 17 | 1784 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT3G02940.1 |
| Entrez Gene | 821178 |
| iHOP | AT3G02940 |
| wikigenes | AT3G02940 |



