 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT32010 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1095aa MW: 122584 Da PI: 5.7619 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 175.3 | 7.9e-55 | 110 | 226 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywlL 102
ke ++rwl+++ei++iL+n+++++++ e+++ p sgsl+L++rk++r+frkDG++w+kkkdgktv+E+he+LK g+++vl+cyYah+een +fqrr+yw+L
EMT32010 110 KEaQTRWLRPTEICEILKNYRNFRIAPEPPNMPASGSLFLFDRKVLRFFRKDGHNWRKKKDGKTVKEAHERLKSGSIDVLHCYYAHGEENINFQRRSYWML 210
5559************************************************************************************************* PP
CG-1 103 eeelekivlvhylevk 118
ee++ +ivlvhylevk
EMT32010 211 EEDYMHIVLVHYLEVK 226
*************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 80.639 | 105 | 231 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.3E-74 | 108 | 226 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 4.5E-48 | 111 | 225 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 6.86E-5 | 523 | 581 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 1.13E-16 | 697 | 801 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.9E-17 | 700 | 803 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 5.69E-13 | 700 | 798 | No hit | No description |
| PROSITE profile | PS50297 | 17.21 | 706 | 800 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.736 | 739 | 771 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.013 | 739 | 768 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1500 | 778 | 807 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 2.92E-8 | 909 | 963 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.036 | 912 | 934 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.498 | 913 | 942 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0036 | 914 | 933 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0022 | 935 | 957 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.578 | 936 | 960 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.1E-4 | 938 | 957 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1095 aa Download sequence Send to blast |
MAGSGSESFT SRSVNSELIS SDPEEEMAAW LALRRSWEEA CERQRSDSFR RESIASAQMA 60 HGSAGGLRVP GGIVGTCKNR SMQMQGHRQG DTRPEYSIHA KLTDMEQILK EAQTRWLRPT 120 EICEILKNYR NFRIAPEPPN MPASGSLFLF DRKVLRFFRK DGHNWRKKKD GKTVKEAHER 180 LKSGSIDVLH CYYAHGEENI NFQRRSYWML EEDYMHIVLV HYLEVKAGKS SSRTRGHDNM 240 LQGAYVDSPL SHLPSQSTDG ESSLSGRASE YEAESGNHQG LQATSPNTGF YSHYQDNSPV 300 IHNESTHGIT FNGPSTQFDL SSWNEMTKMN KGIHQLPPYQ SHVPSEQRPF TEGPGIESFS 360 FDEVYSNGLD IKDDGHADTD REALWQLPSA NDGTTTEFLQ LPSAIDGRTT EFQLPSATDS 420 TFATVDSFEQ NNKLLEEAIN FPVLKTQSSN LSDILKNSFK KSDSFTRWMS KELAEVDDSQ 480 VKSSSALYWN SEDADNIIGA SSRDQLDQFT LDPMVAQDQL FSITEYFPSW TYAGSKTRVL 540 VTGRFLTSDE VIKLKWSCMF GEVEVPADIL ADGTLRLACS EVREFEYRPS DSQYMDAPSP 600 HGATNKIYLQ ARLDELLSLG QDEQDEFQAA LSNPTKELID LNKKITSLMT NNDQWSELLK 660 FADDNQLAPD DRQDQFVESG IKEKLHIWLL HKAGGGGKGP SVLDDEGQGV LHLAAALGYD 720 WAIRPTITAG VSINFRDVHG WTALHWAAFC GRERTVVALI ALGAAPGALT DPRPDFPSGR 780 TPADLASFNG HKGISGFLAE FSLTSHLQTL NLKEAMGSNA SEISGLPGIG DVTGRIASPS 840 AGQGLQAGSM GDSLGAVRNA AQAAARIYQV FRVQSFQRKQ AVQYEDDNGV ISDERAMSLL 900 SYKPSKPGQF DPMHAAATRI QNKFRGWKGR KEFLLIRQRI VKIQAHVRGH QVRKHYRKII 960 WSVGIVEKVI LRWRRRGAGL RGFRSTEVAT DSSTSSSSVD VIPVKPAEDD YNFLQEGRKQ 1020 TEERLQRALA RVKSMVQYPE ARDQYQRILT VVTKMQESQP VEESMLEEST EMDEGFLMSE 1080 FKELWDDDVP LPGYF |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
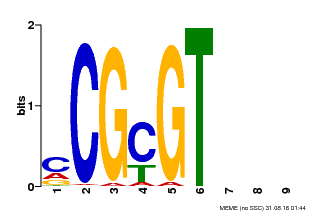
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK365210 | 0.0 | AK365210.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2032B14. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020179696.1 | 0.0 | calmodulin-binding transcription activator 3-like isoform X2 | ||||
| TrEMBL | M8C3X6 | 0.0 | M8C3X6_AEGTA; Calmodulin-binding transcription activator 3 | ||||
| STRING | EMT32010 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



