 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT20356 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 579aa MW: 64633.1 Da PI: 8.071 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 18.6 | 5.3e-06 | 138 | 160 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ Cg +F+ + +Lk+H+++H
EMT20356 138 HDCKECGARFKKPAHLKQHMQSH 160
56*******************99 PP
| |||||||
| 2 | zf-C2H2 | 18.8 | 4.3e-06 | 296 | 320 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ C+ C+ s+srk++L+rH+ tH
EMT20356 296 FACHvdGCPFSYSRKDHLNRHLLTH 320
78*********************99 PP
| |||||||
| 3 | zf-C2H2 | 10.7 | 0.0017 | 325 | 350 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+ Cp C++ Fs k n +rH H
EMT20356 325 FMCPmeGCNRKFSIKGNIQRHVEEfH 350
78*******************98767 PP
| |||||||
| 4 | zf-C2H2 | 19.7 | 2.2e-06 | 362 | 386 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp +Cgk+F+ s L++H +H
EMT20356 362 FICPeaNCGKVFKYASKLQKHEESH 386
78*******************9988 PP
| |||||||
| 5 | zf-C2H2 | 20.4 | 1.4e-06 | 453 | 477 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ dC++sFs ksnL++Hi+ H
EMT20356 453 KCNfkDCKRSFSKKSNLHKHIKAvH 477
79999****************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 11 | 37 | 59 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 3.6E-6 | 137 | 174 | No hit | No description |
| PROSITE profile | PS50157 | 12.279 | 138 | 165 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-5 | 138 | 162 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0037 | 138 | 160 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 140 | 160 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 6.35E-7 | 291 | 322 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.8E-9 | 292 | 322 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0018 | 296 | 320 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.307 | 296 | 325 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 298 | 320 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.28E-6 | 323 | 351 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-6 | 323 | 351 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.245 | 325 | 355 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.029 | 325 | 350 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 327 | 350 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.84E-5 | 358 | 388 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.0E-7 | 358 | 386 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0022 | 362 | 386 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.115 | 362 | 391 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 364 | 386 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 6.5 | 394 | 419 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 396 | 419 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 12 | 422 | 443 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 3.04E-8 | 433 | 490 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.5E-6 | 433 | 474 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0048 | 452 | 477 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.36 | 452 | 482 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 454 | 477 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 5.8E-4 | 476 | 505 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 8.6 | 483 | 509 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 4.8 | 483 | 509 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 485 | 509 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 579 aa Download sequence Send to blast |
MGEGDPDSGG GVGATSASPS HAPRPAAAAP VRDIRRYKCE FCDVVRSKKR LIRDHVLEHH 60 KSTRATDFCG SCELWVSLGL DLDLAAAGGG RPCEGHQAST RATDFCGSCE LWDEVDGLDE 120 YNVGGGGGSA PPGKEIGHDC KECGARFKKP AHLKQHMQSH SPESTRATDF CGSCELWDEV 180 DGLDEYNVGG GGGSAPPGKE IGHDCKECGA RLFNCVKHWI PKNSKMREGP EWTSTRMGRP 240 SWEAPSGSLF HALLQHWIPR TPKCQKALNG PVSGRGDPPE KLHQECLIHA LPQRPFACHV 300 DGCPFSYSRK DHLNRHLLTH QGKLFMCPME GCNRKFSIKG NIQRHVEEFH EDGPQCGGKK 360 EFICPEANCG KVFKYASKLQ KHEESHVNLD YTEVICCEPG CMKTFTNVEC LKAHNQSCHQ 420 YVQCDICDTK QLKKNFKRHQ RMHEGSFVTE RIKCNFKDCK RSFSKKSNLH KHIKAVHEQS 480 RPFTCGFSGC GQKFSYKHVR DNHEKSSVHV LFEELIIRTL FTSWQGDFVE ADEQLRPHPG 540 GRKRKPISVD TFMRKRVAAP DAPPACSDGT EYLRWLLSG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2i13_A | 6e-13 | 293 | 506 | 19 | 183 | Aart |
| 2i13_B | 6e-13 | 293 | 506 | 19 | 183 | Aart |
| Search in ModeBase | ||||||
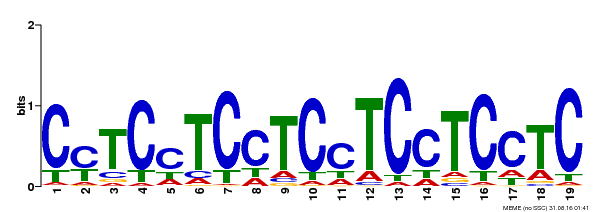
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK372817 | 0.0 | AK372817.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, partial cds, clone: NIASHv3013K06. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020149307.1 | 0.0 | transcription factor IIIA-like | ||||
| Refseq | XP_020149308.1 | 0.0 | transcription factor IIIA-like | ||||
| Refseq | XP_020149309.1 | 0.0 | transcription factor IIIA-like | ||||
| Refseq | XP_020149310.1 | 0.0 | transcription factor IIIA-like | ||||
| TrEMBL | R7WCP6 | 0.0 | R7WCP6_AEGTA; Transcription factor IIIA | ||||
| STRING | EMT20356 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 2e-82 | transcription factor IIIA | ||||



