 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT19288 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1121aa MW: 123771 Da PI: 8.2599 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.5 | 0.00045 | 999 | 1021 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L +H ++H
EMT19288 999 CPvnGCGKKFFTHKYLLQHRKVH 1021
9999*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51184 | 34.108 | 1 | 162 | IPR003347 | JmjC domain |
| SMART | SM00558 | 2.4E-41 | 1 | 162 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 5.49E-25 | 9 | 159 | No hit | No description |
| Pfam | PF02373 | 8.6E-37 | 26 | 145 | IPR003347 | JmjC domain |
| SMART | SM00355 | 17 | 900 | 922 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 1.8 | 923 | 947 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.847 | 923 | 952 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 4.5E-4 | 924 | 948 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 925 | 947 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 46 | 974 | 996 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.56 | 997 | 1021 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.487 | 997 | 1026 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 999 | 1021 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 6.6E-6 | 999 | 1020 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.55E-14 | 1008 | 1064 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.5E-7 | 1021 | 1047 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0062 | 1027 | 1051 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.323 | 1027 | 1056 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1029 | 1051 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.1E-7 | 1048 | 1069 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0032259 | Biological Process | methylation | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0008168 | Molecular Function | methyltransferase activity | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1121 aa Download sequence Send to blast |
MRAAARGPGS LLRFMRDEVP GVTTPMLYVG MLFSWFAWHV EDHDLHSLNY MHLGAAKTWY 60 GVPRDAALAF EDVVRVHGYG GEVNALEAFA TLGQKTTLLS PELLVGSGVP CCRLVQNAGE 120 FVVTFPGSYH CGFSHGFNCG EASNIATPEW LRVAKEAAVR RASINQPPLL SHYQLLYELA 180 LSMCIRDPSI GAMEPRSSRL KEKKKGEGGQ LVKKLFVQNA IQDNELLSCL LNDGSSCIIL 240 PINAHDGPVL SALRSRSQLI PKSKLSRCTH DSHNAEGDKA DVISAAGLLD QGLLSCVSCG 300 ILSFSCVAVI KPRECASKYF MSSDYNSIND QIVGSGGSHL ANAAGSEGTN GGILRPSFEP 360 YGNAILPDAG PVTRNSAPDL LASPYGNQPD TDEDNRNKKI KVSHDSSELD GTKIPSSSIK 420 CQQRPSSQSS QCIGGSSISN GPRGVRTRNK YRLKMALSQG FQLKNNYWTM EQKVQPEPSR 480 SKETVKEPLD VNGAENDATC NSAAISVGDP RISTTTMDNL NKPIVKIDKD SSRMHVFCLE 540 HAVEVEKQLQ AIGGAHVILL CRPEYLKIEA EARSLAAELE VEHGWKDIHF REANMEDRKM 600 IEELLQDEES IPTSSDWAVK LGTNLYYSAN LAKSPLHNKQ IPYNRVIYRA FGCSSPDNSP 660 VKLENCEGSQ DRQKKIVLAG RWCGKAWMSN QVHPFLAQRI ETSELEEADK SSGVEASKRK 720 SSTITNVPKS SKKRENMAVE VTTDTKRPRL AEGHSSKALK GVAEVSHPAP TAVVPRVSSR 780 IADRANKLKS EMTEEDDDPA CRPKPKVTSH SRRRPPKKIE VEAKKQMKMS KGDKMMVPTA 840 PNDDEEHPSA AKGGPSAGPA TKLELSPRKH RTRTKTMKQL KKATGEERTP RDHPMHVEGY 900 TCSIEGCSMS FDTRNELSLH ELDICPVKGC GKKFFTHKFL LQHRKVHTET MKQLKKATGE 960 ERAPRDHPMR VEGYTCGIES CSMSFDTKKE LSLHERDICP VNGCGKKFFT HKYLLQHRKV 1020 HTEDRPLKCP WEGCDVAFKW AWARTEHLRV HTGDRPYVCR EPGCAETFSC ELVGGGSAQK 1080 KGTDPLAVVD PFVLLGCPPS LWDGMLALLF GAVRIETWEW D |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 2e-51 | 1 | 183 | 167 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 2e-51 | 1 | 183 | 167 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3 with a specific activity for H3K27me3 and H3K27me2. No activity on H3K4me3, H3K9me3, H3K27me1 and H3K36me3. Involved in biotic stress response. May demethylate H3K27me3-marked defense-related genes and increase their basal and induced expression levels during pathogen infection. {ECO:0000269|PubMed:24280387}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
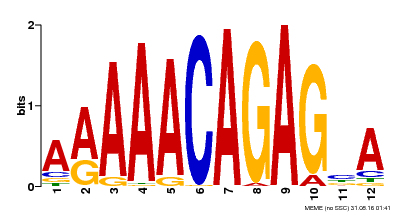
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt stress, abscisic acid (ABA), jasmonic acid (JA), the ethylene precursor ACC and infection by the bacterial pathogen Xanthomonas oryzae pv. oryzae. {ECO:0000269|PubMed:24280387}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG670306 | 0.0 | HG670306.1 Triticum aestivum chromosome 3B, genomic scaffold, cultivar Chinese Spring. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020164795.1 | 0.0 | lysine-specific demethylase JMJ705-like | ||||
| Swissprot | Q5N712 | 0.0 | JM705_ORYSJ; Lysine-specific demethylase JMJ705 | ||||
| TrEMBL | R7WBU2 | 0.0 | R7WBU2_AEGTA; Lysine-specific demethylase 5A | ||||
| STRING | EMT19288 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5346 | 32 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 1e-122 | relative of early flowering 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



