 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT09464 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 569aa MW: 62929 Da PI: 6.171 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 28.6 | 3.1e-09 | 212 | 264 | 11 | 63 |
HHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 11 qkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
NR +A rs +RK +++ eLe+kv++L++e ++L +l l+ + l++e+
EMT09464 212 LANRQSAARSKERKIKYTGELERKVQTLQTEATTLSAQLTLLQRDTSGLTTEN 264
56***************************************999888888776 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 4.9E-12 | 202 | 266 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14703 | 7.25E-22 | 208 | 256 | No hit | No description |
| SuperFamily | SSF57959 | 2.46E-9 | 209 | 257 | No hit | No description |
| Pfam | PF00170 | 2.0E-7 | 210 | 264 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 5.5E-10 | 210 | 261 | No hit | No description |
| PROSITE profile | PS50217 | 9.588 | 210 | 267 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007231 | Biological Process | osmosensory signaling pathway | ||||
| GO:0008272 | Biological Process | sulfate transport | ||||
| GO:0009294 | Biological Process | DNA mediated transformation | ||||
| GO:0009410 | Biological Process | response to xenobiotic stimulus | ||||
| GO:0009652 | Biological Process | thigmotropism | ||||
| GO:0009970 | Biological Process | cellular response to sulfate starvation | ||||
| GO:0045596 | Biological Process | negative regulation of cell differentiation | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0051170 | Biological Process | nuclear import | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0043621 | Molecular Function | protein self-association | ||||
| GO:0051019 | Molecular Function | mitogen-activated protein kinase binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 569 aa Download sequence Send to blast |
MDPRFQLPTP VPSSGGAGVR GHHRRAHSET FLRFPDTDLF LDPDGDFSFS DLDFPSLSDD 60 SPAVSDPTPP PPPPMAASSS QTPAPRPPGG THNRSLSLDA AFFEGLALQG GGGGGCGGHK 120 RSGSMDGVNS PFEGESALSG GLDYAKKAMP AERIAELALL DPKRAKSSEI LDIWMIVARS 180 RVCLLVVFGL YEFRIIVSFE ISDIWMIVAR ILANRQSAAR SKERKIKYTG ELERKVQTLQ 240 TEATTLSAQL TLLQRDTSGL TTENRELKLR LQSMEEQAKL RDALNDALRE EVQRLKIAAG 300 QAPNMNGSQF NGGLQQIPSY FSQQHQRQQQ QQHQQQMAYL GGHQAQRHHP NHHQSPSNGG 360 QSLSGHNCYE IHLLDTELCT CSSLLASLEL LRGCFKSNLF ILLFEVYIPF FVCFWRAIDD 420 TGNSYLQCAF QVGSSKIDVL KYIHGLTLAT GPRALWMSWA QYRSPQVQPL ANCPLLAALP 480 GDQGEDEQRQ LRDATAMDPV VGQEETSFAI PDLRLARGTH NRHDLGIRRQ REVDEADGAE 540 IEEGVVINTL WEQGDVRLIK LLESTVLRE |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00184 | DAP | Transfer from AT1G43700 | Download |
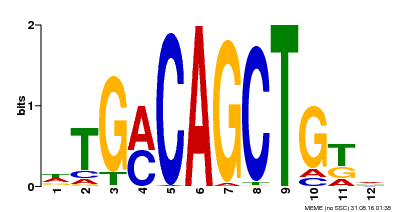
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK372616 | 0.0 | AK372616.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv3007C09. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020172126.1 | 0.0 | transcription factor VIP1-like | ||||
| TrEMBL | M8BRQ5 | 0.0 | M8BRQ5_AEGTA; Putative transcription factor PosF21 | ||||
| STRING | EMT09464 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1925 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G43700.1 | 4e-40 | VIRE2-interacting protein 1 | ||||



