 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 883868 | ||||||||
| Common Name | ARALYDRAFT_683312 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 504aa MW: 56454.3 Da PI: 4.9723 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 37.5 | 4.2e-12 | 337 | 382 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksL 54
+h e+Er RR+++N++f Lr ++P+ + K++K + Le Av+YI++L
883868 337 NHVEAERMRREKLNHRFYALRAVVPNiS------KMDKTSLLEDAVHYINEL 382
799***********************66......***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 2.6E-49 | 34 | 228 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| CDD | cd00083 | 1.44E-13 | 332 | 383 | No hit | No description |
| PROSITE profile | PS50888 | 16.463 | 333 | 382 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 5.37E-17 | 333 | 404 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.6E-9 | 337 | 382 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.5E-15 | 337 | 404 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.4E-14 | 339 | 388 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005509 | Molecular Function | calcium ion binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 504 aa Download sequence Send to blast |
MINSDDNLSM IEALFTSDLS PLPPANLSLE TTLPKRLHAV LNGTNEPWTY VIFWKPSYDY 60 DISGESVLKW SDGVYNGGDE EKTRERLRRK KTIPSSPAER ERRSNVLREL NSMISGEAFP 120 VVEDEYVNKD DDVEAEVTDM EWFFLVSMTW SFGSGSGLAG KAFASYNPVW VTGSDQIYGS 180 GCDRAKQGGD LGLQTIVCIP SDNGVLELGS TEHIQQNSDL FNRIRFLFNF DGSKDFPGAP 240 NLNSELFSFQ LETGFSSTVT DNPNPSYNLN FSTSCSTSAR ASCGDVLSFS DIVKQSSENL 300 NPNTYSDQIQ NATVMPEKKQ GKKRGRKPAH GRDQPLNHVE AERMRREKLN HRFYALRAVV 360 PNISKMDKTS LLEDAVHYIN ELKSKAENAE SEKNAIQIQL NELKEMAGQR NAIPSVFKYE 420 ENASEMKIEV KIMGNDAMVR VESSKSHHPG ARLMNALMDL ELEVNNASMS VMNDFMIQQA 480 NVKMGLRIYK QEELRDVLIS KIS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rqw_A | 1e-47 | 30 | 231 | 6 | 195 | Transcription factor MYC3 |
| 4rqw_B | 1e-47 | 30 | 231 | 6 | 195 | Transcription factor MYC3 |
| 4rs9_A | 1e-47 | 30 | 231 | 6 | 195 | Transcription factor MYC3 |
| 4yz6_A | 1e-47 | 30 | 231 | 6 | 195 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00547 | DAP | Transfer from AT5G46830 | Download |
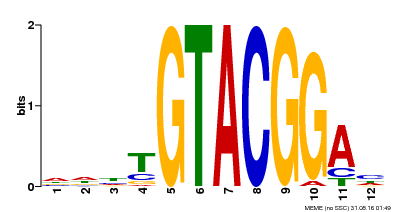
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 883868 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By UV treatment. {ECO:0000269|PubMed:12679534}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB022221 | 0.0 | AB022221.1 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MSD23. | |||
| GenBank | AB493781 | 0.0 | AB493781.1 Arabidopsis thaliana At5g46830 mRNA for hypothetical protein, partial cds, clone: RAAt5g46830. | |||
| GenBank | AF252636 | 0.0 | AF252636.1 Arabidopsis thaliana putative transcription factor bHLH28 mRNA, complete cds. | |||
| GenBank | CP002688 | 0.0 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002865159.2 | 0.0 | transcription factor bHLH28 | ||||
| Swissprot | Q9LUK7 | 0.0 | BH028_ARATH; Transcription factor bHLH28 | ||||
| TrEMBL | D7MQN9 | 0.0 | D7MQN9_ARALL; Predicted protein | ||||
| STRING | Al_scaffold_0008_132 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM13953 | 16 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G46830.1 | 0.0 | NACL-inducible gene 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 883868 |
| Entrez Gene | 9301233 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



