 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 491407 | ||||||||
| Common Name | ARALYDRAFT_491407 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 310aa MW: 34452.3 Da PI: 4.9289 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 54.5 | 1.6e-17 | 200 | 234 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C +C t kTp+WR gp g+ktLCnaCG++y++ +l
491407 200 CLHCATEKTPQWRTGPMGPKTLCNACGVRYKSGRL 234
99*****************************9885 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF016992 | 2.9E-110 | 1 | 290 | IPR016679 | Transcription factor, GATA, plant |
| SMART | SM00401 | 1.7E-17 | 194 | 244 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 12.242 | 194 | 230 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 1.43E-15 | 196 | 258 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 2.2E-14 | 197 | 232 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 1.60E-14 | 199 | 246 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 200 | 225 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 2.6E-15 | 200 | 234 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 310 aa Download sequence Send to blast |
MERLTSELFL VAGNTDSFVV DDLLDFSNDD GEIDDGFDTL PDSSALSTGT LTDSSNSSSL 60 FTDGTGFSDL CVPRDDIAEL EWLSNFVEES FSGEVQDKLH LLSGLKNPQT TGSTLTHLIK 120 PEPEPDFDQF IDIDESNVAV PAKARSKRSR SAASTWASRL LSLADSNETN PKKKQRRVKE 180 QDFAADMDVD CGETGGGRRC LHCATEKTPQ WRTGPMGPKT LCNACGVRYK SGRLVPEYRP 240 ASSPTFVMAR HSNSHRKVME LRRQKEMRDE HLLSQLRCEN LLMDIRSNGE DLVMHNNNNH 300 VAPDFRHLI* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 170 | 176 | PKKKQRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. May be involved in the regulation of some light-responsive genes (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00048 | PBM | Transfer from AT4G32890 | Download |
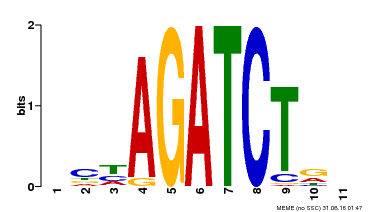
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 491407 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK117169 | 0.0 | AK117169.1 Arabidopsis thaliana At4g32890 mRNA for unknown protein, complete cds, clone: RAFL16-71-E16. | |||
| GenBank | BT008342 | 0.0 | BT008342.1 Arabidopsis thaliana At4g32890 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020872057.1 | 0.0 | GATA transcription factor 9 | ||||
| Swissprot | O82632 | 0.0 | GATA9_ARATH; GATA transcription factor 9 | ||||
| TrEMBL | D7M9N0 | 0.0 | D7M9N0_ARALL; GATA transcription factor | ||||
| STRING | fgenesh2_kg.7__840__AT4G32890.1 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5682 | 27 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G32890.1 | 0.0 | GATA transcription factor 9 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 491407 |
| Entrez Gene | 9303294 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



