 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araip.2D8LN | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Dalbergieae; Arachis
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 403aa MW: 46010.9 Da PI: 4.5923 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 109.3 | 2.8e-34 | 14 | 105 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkellekik 98
Fl k+ye+++d++++ ++sws++++sf+v+++ +fa+++Lp++Fkh+nf+SF+RQLn+YgFkkv+ e+ weF+++ F +g+ + +++i+
Araip.2D8LN 14 FLAKTYEMVDDPSTDPIVSWSASNRSFIVWNPPDFARDLLPRFFKHNNFSSFIRQLNTYGFKKVDPEQ---------WEFANDDFVRGQPHRMKNIH 101
9********************999****************************************9998.........******************** PP
XXXX CS
HSF_DNA-bind 99 rkks 102
r+k
Araip.2D8LN 102 RRKP 105
*985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 9.8E-38 | 3 | 97 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SuperFamily | SSF46785 | 7.62E-35 | 9 | 103 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 1.1E-61 | 10 | 103 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.8E-19 | 14 | 37 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 1.8E-30 | 14 | 103 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.8E-19 | 52 | 64 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 53 | 77 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.8E-19 | 65 | 77 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 403 aa Download sequence Send to blast |
MDEAQSSSNS LPPFLAKTYE MVDDPSTDPI VSWSASNRSF IVWNPPDFAR DLLPRFFKHN 60 NFSSFIRQLN TYGFKKVDPE QWEFANDDFV RGQPHRMKNI HRRKPVHSHS LQNLQAQAPL 120 TESERMSLQD EIERLNREKE QLLMELQRHE QEWQEYEIRI HCSKDRVEKM EQKHQKMISS 180 VSSVLQKPGI DLNQLALLET TDRKRRLPRN GYLSDETNIE DTTETEQMLP TENSQDSSVL 240 TIITEQLDQL ESSMVFWETI SNDISDATIH IHSSLDFDES TSCADSLSIS CPLLDVEVLP 300 KSSGIDMNSE PASAPVSEPA SASKEQPAGT TPVVAGVNDV FWEQFLTENP GSSEAQEVQS 360 ERKYHDTSKG EGKPSDPGKF WWNMRNTSNL PEQIGHVSQA EKT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 3e-23 | 6 | 103 | 21 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 3e-23 | 6 | 103 | 21 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 3e-23 | 6 | 103 | 21 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 201 | 206 | DRKRRL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds DNA sequence 5'-AGAAnnTTCT-3' known as heat shock promoter elements (HSE). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
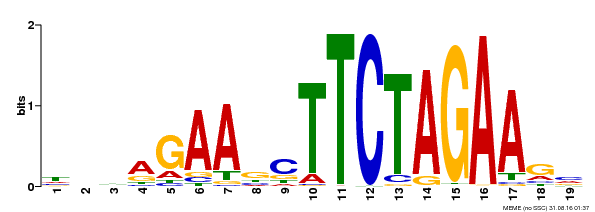
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araip.2D8LN |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016191319.1 | 0.0 | heat stress transcription factor A-4c | ||||
| Refseq | XP_025640334.1 | 0.0 | heat stress transcription factor A-4c | ||||
| Swissprot | O49403 | 1e-119 | HFA4A_ARATH; Heat stress transcription factor A-4a | ||||
| TrEMBL | A0A444XBG1 | 0.0 | A0A444XBG1_ARAHY; Uncharacterized protein | ||||
| STRING | GLYMA08G12630.3 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3568 | 33 | 61 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G45710.1 | 1e-115 | HSF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



