 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_7729_f_3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 402aa MW: 42872.9 Da PI: 9.7358 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 45.8 | 1.3e-14 | 312 | 355 | 5 | 48 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkke 48
+r rr++kNRe+A rsR+RK+a++ eLe +v++L++ N++L+k+
Neem_7729_f_3 312 RRHRRMIKNRESAARSRARKQAYTLELEAEVAKLKELNQELQKK 355
699************************************99976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 7.7E-11 | 308 | 370 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.415 | 310 | 355 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 1.3E-12 | 312 | 355 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 7.03E-10 | 312 | 356 | No hit | No description |
| CDD | cd14707 | 3.07E-24 | 312 | 357 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 9.9E-14 | 312 | 356 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 315 | 330 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 402 aa Download sequence Send to blast |
MGSQMNFKNF SDTLHGEGKP PGNTPLARQP SVYSLTFEEF QNTWGGLGKD FGSMNMDELL 60 KNIWTAEESQ AITSSAGATG EGNVPGGNLQ RQGSLTLPRT LSQKTVDEVW RDLIKESSGG 120 TKDGSNGSGV SGGGAGANLP QRQQTLGEMT LEEFLVRAGV VREDTQQIGG RLNNGGFYAN 180 NNNTSLALGF QQPNRNNGLM GNRVMQDSNS VPRQPPSLTL NLTSPVARGP VVTVANPSMN 240 NNLVQAAGLQ GGGMGVVGLG AAGITIATGS PAGQVSPDVI AKSTVDTSSP SPVPYAIGRG 300 RRSSALEKVV ERRHRRMIKN RESAARSRAR KQAYTLELEA EVAKLKELNQ ELQKKQVSFL 360 YQEERVEIHR NEKRVFEHAI LAADAVQGSG CASVIHISSY GV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
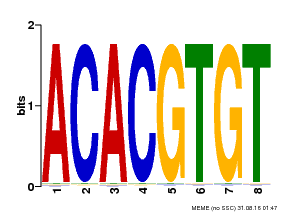
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022763380.1 | 1e-150 | ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_022763381.1 | 1e-150 | ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_022763382.1 | 1e-150 | ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Swissprot | Q9M7Q3 | 2e-99 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | A0A3G2FPB6 | 1e-163 | A0A3G2FPB6_LITCN; ABF1 | ||||
| STRING | POPTR_0009s10400.1 | 1e-147 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM734 | 27 | 129 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G45249.1 | 3e-57 | abscisic acid responsive elements-binding factor 2 | ||||



