 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_6747_f_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 593aa MW: 64501.8 Da PI: 9.4792 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.2 | 3.1e-05 | 52 | 74 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ C+k F r nL+ H r H
Neem_6747_f_1 52 FICEVCNKGFQREQNLQLHRRGH 74
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 13.1 | 0.00028 | 128 | 150 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC++C+k++ +s+ k H +t+
Neem_6747_f_1 128 WKCEKCSKRYAVQSDWKAHSKTC 150
58*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF12171 | 2.2E-5 | 51 | 74 | IPR022755 | Zinc finger, double-stranded RNA binding |
| SMART | SM00355 | 0.0052 | 52 | 74 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.5E-5 | 52 | 74 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.99 | 52 | 74 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.43E-7 | 52 | 74 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 54 | 74 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 130 | 93 | 123 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-5 | 116 | 149 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.43E-7 | 123 | 148 | No hit | No description |
| SMART | SM00355 | 130 | 128 | 148 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 593 aa Download sequence Send to blast |
MSASSSAPFF EIREEDQNQM KQQHSATPTS SSAPAPPPQK KKRNQPGTPS KFICEVCNKG 60 FQREQNLQLH RRGHNLPWKL KQKTTKEVKR KVYLCPEPTC VHHDPSRALG DLTGIKKHYS 120 RKHGEKKWKC EKCSKRYAVQ SDWKAHSKTC GTREYRCDCG TLFSRRDSFI THRAFCDALA 180 QESARQPPSL SAISSHLYGS SNMGLGLSQV GPQLSSIKDQ NSQSNDILRL GGGSRSTAFD 240 HLLPPSMGSS SSFRPPQNMA STGFFMQESN QNYHEEHPSQ QGLLPNKTFH GLMQFPDLQN 300 NTSNTPSAAN LFNLSFLSNS SSSSSISNSN NANTTNSNLQ SSNLLISGHF NSDQNGTTTG 360 GVEGSNLFSN NLINDQISSG VPSLFSSSVQ NENMVPHMSA TALLQKAAQM GSTSSNNNTA 420 SLLRSFGSSS SGGTKPNFGG IFGAPENDSN LQDLMNTFAS GNSSIFGTGS VEVNAYNSPH 480 GQENPYAGYN TNRTNLEQEK RNHGQSFGNI DERKLHQNLN ISNIGGSDRM TRDFLGVGQI 540 VRSVSGGFSQ REKQQQQQQH RHGIDVSSLD SERNVAAPTS QSFGGAGGGG SFQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 9e-34 | 124 | 185 | 3 | 64 | Zinc finger protein JACKDAW |
| 5b3h_F | 9e-34 | 124 | 185 | 3 | 64 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a positive regulator of the starch synthase SS4. Controls chloroplast development and starch granule formation (PubMed:22898356). Binds DNA via its zinc fingers (PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). {ECO:0000269|PubMed:22898356, ECO:0000269|PubMed:24821766}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
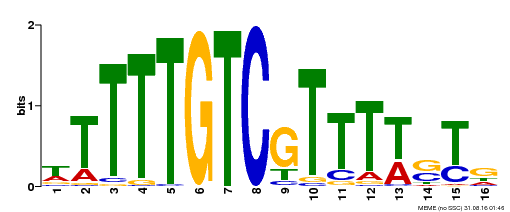
| Motif ID | Method | Source | Motif file |
| MP00255 | DAP | Transfer from AT2G02070 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC208494 | 1e-137 | AC208494.1 Populus trichocarpa clone JGIACSB265-L05, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011014313.1 | 0.0 | PREDICTED: zinc finger protein NUTCRACKER-like | ||||
| Refseq | XP_011014314.1 | 0.0 | PREDICTED: zinc finger protein NUTCRACKER-like | ||||
| Refseq | XP_011014315.1 | 0.0 | PREDICTED: zinc finger protein NUTCRACKER-like | ||||
| Swissprot | Q9ZUL3 | 1e-141 | IDD5_ARATH; Protein indeterminate-domain 5, chloroplastic | ||||
| TrEMBL | A0A2K1ZH36 | 0.0 | A0A2K1ZH36_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0008s14180.4 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4373 | 26 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G02070.1 | 1e-102 | indeterminate(ID)-domain 5 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



