 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_26782_f_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 609aa MW: 67582.8 Da PI: 6.2255 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 40.5 | 5e-13 | 440 | 486 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI++Lq
Neem_26782_f_1 440 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAIAYINELQ 486
799***********************66......******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 2.2E-56 | 46 | 234 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| SuperFamily | SSF47459 | 3.27E-18 | 435 | 499 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.047 | 436 | 485 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 8.36E-15 | 439 | 490 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 2.9E-18 | 440 | 500 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.8E-10 | 440 | 486 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.8E-16 | 442 | 491 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 609 aa Download sequence Send to blast |
MGSNLWNDED KAMVAAVLGT RAFDYLRSSS ISNENLLMAV GSDENLQNKL SDLVDRPNAS 60 NFSWNYAIFW QISRSKSGDW VLGWGDGSCR EPKEGEESEA TRILSIRLED ETEQKMRKRV 120 LQKLHTLFGG SDEDNYALGL DRVTDTEMFF LASMYFSFPR GEGGPGKCFA SGKHVWLLDT 180 LQVGSDYCVR SFLAKSAGIQ TIVLITTDVG VVELGSVRSV PESLELVQSI RAMFSSNPSV 240 TRVKPMTAVL PIVSEKKDEN ALFSNLGILD RVEAVPKIFG QDLNNSGRIQ PHFSREKLAV 300 RKTEDRPSWE AYANGSRTPF PSVRNGVHGF SWGNVQAVKQ GSPTEIFGSQ TTSNNLQDLV 360 NGVREDFRVN HYQSQKPVQM QIDFSGATSR PTVIARPLSA DSEHSDVEAS CRDERPSVAD 420 ERRPRKRGRK PANGREEPLN HVEAERQRRE KLNQRFYALR AVVPNISKMD KASLLGDAIA 480 YINELQAKLK VMEAEREKFS STSRDSSALE TIPNAESQNP SPDVDIQAAH DEVVVRVSCP 540 LDSHPASRVI QAFKESQITV VESKLSAVND TVFHTFVIKS QGSEQLTKEK LIAAFSCESN 600 SLQPLSSVG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 1e-29 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 1e-29 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 1e-29 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 1e-29 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 1e-29 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 1e-29 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 1e-29 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 1e-29 | 434 | 496 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 421 | 429 | RRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
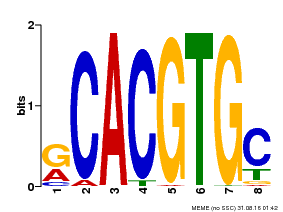
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006444826.1 | 0.0 | transcription factor bHLH13 | ||||
| Swissprot | A0A3Q7ELQ2 | 0.0 | MTB1_SOLLC; Transcription factor MTB1 | ||||
| TrEMBL | A0A2H5P035 | 0.0 | A0A2H5P035_CITUN; Uncharacterized protein | ||||
| TrEMBL | V4U7A9 | 0.0 | V4U7A9_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006444826.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3758 | 27 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 0.0 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



