 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_2178_f_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 492aa MW: 54505.2 Da PI: 7.0082 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.3 | 3.1e-17 | 60 | 104 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE++++v+a k++G W++I +++g ++t+ q++s+ qk+
Neem_2178_f_1 60 RERWTEEEHKKFVEALKLYGRA-WRKIEEHVG-TKTAVQIRSHAQKF 104
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 7.48E-17 | 54 | 108 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.908 | 55 | 109 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-9 | 57 | 104 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.7E-17 | 58 | 107 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.0E-13 | 59 | 107 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.5E-14 | 60 | 103 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.69E-11 | 62 | 105 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 492 aa Download sequence Send to blast |
MAVQDQCGGT HSNSAFSTGN EISYDAGLHP VTGLEIKDQF SCGNDYAPKP RKPYTITKQR 60 ERWTEEEHKK FVEALKLYGR AWRKIEEHVG TKTAVQIRSH AQKFFTKVAR DCSGYSTSPM 120 EPIDIPPPRP KRKPMHPYPR KLVYPRVRES PNPELPRRSS SPNLSTSEPE NQSPTSVFSA 180 VGSDAFGSSD SNSPNDSLSP VSPVAPAHLA GFTVSQPNSS PAENIPPSPV LETTDSVPDE 240 QSPMDVLKLG LSSQEFAKEG VPTRTLKLFG TTVLVTDSQR PMSSTPGACK LSPQHVNEGK 300 PLQLIPLKLS ESPAGDTKCF QKQLLHTGSE APCCIHCPDE NLSLIEGGSG ARLPRWTFCG 360 GILFPITPYG QPEPMKMHLD SILKEVQDKE FWKEGSWTGS NSKSINEDDN FDKNSDVETQ 420 SCQQPLDKEE KGSDKVFELK PSENSAFSVI RSSPMKHIKG FVPYKKRMAE RDHQLSAVTG 480 DEREEQRLRH CY |
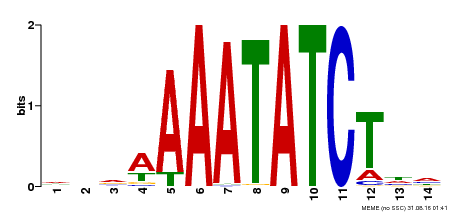
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006481961.1 | 0.0 | protein REVEILLE 1 | ||||
| TrEMBL | A0A2H5MZZ0 | 0.0 | A0A2H5MZZ0_CITUN; Uncharacterized protein | ||||
| STRING | XP_006481961.1 | 0.0 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6356 | 26 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G17300.1 | 9e-58 | MYB_related family protein | ||||



