 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_1329_f_4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 447aa MW: 50892.7 Da PI: 8.2081 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.9 | 8.6e-17 | 203 | 249 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g WT++Ed+ l ++v q+G++ W+ I++++ gR +kqc++rw+++l
Neem_1329_f_4 203 KGQWTPQEDKTLMRLVSQYGTKKWSQISKMLT-GRVGKQCRERWYNHL 249
799****************************9.*************97 PP
| |||||||
| 2 | Myb_DNA-binding | 52.6 | 1e-16 | 255 | 297 | 1 | 45 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwq 45
+++WT++Ed +l++a+k+ G++ W+ Iar++ gRt++ +k++w+
Neem_1329_f_4 255 KDSWTEDEDIILIQAHKEVGNR-WAEIARRLR-GRTENTIKNHWN 297
679*******************.*********.***********8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 26.965 | 198 | 253 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.26E-30 | 201 | 296 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.1E-15 | 202 | 251 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.4E-27 | 204 | 256 | IPR009057 | Homeodomain-like |
| Pfam | PF13921 | 1.1E-18 | 206 | 264 | No hit | No description |
| CDD | cd00167 | 5.03E-14 | 206 | 249 | No hit | No description |
| SMART | SM00717 | 7.0E-16 | 254 | 302 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 21.109 | 254 | 304 | IPR017930 | Myb domain |
| CDD | cd00167 | 1.08E-12 | 257 | 300 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.8E-20 | 257 | 302 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010439 | Biological Process | regulation of glucosinolate biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1904095 | Biological Process | negative regulation of endosperm development | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 447 aa Download sequence Send to blast |
MEFDSSFRDD YPFLASLFSD NLFKSDLENG FSFLENSSSK GLFHNLHTSV DHGPISECFS 60 SPDQFNRFST EGSSKNPSFG ISTPSFDAIE AYTNRVSGSG IGFSSTHLMP NGVNEVLHGS 120 ESGRFWDYSQ NFSAQPASET QIYQPVMNYP QCGSSSTARL PDEVSCITGE NAYHQDQKAD 180 QKRCNRMHVR KGRKLPKRNS IIKGQWTPQE DKTLMRLVSQ YGTKKWSQIS KMLTGRVGKQ 240 CRERWYNHLK PDIKKDSWTE DEDIILIQAH KEVGNRWAEI ARRLRGRTEN TIKNHWNATK 300 RRQQSKRKHK DRTPRSNLLQ NYIKTVTSPP STRKKKGKAH AASNISDQVV KTLPPTPQTQ 360 QVEISSDFTP SEWQVPAYDP NDATGFSYET SMLSDGYIFR SALDEVPSGS IVESNLDFEI 420 PTEIDSNYMP EEVTREMDLM VMVYKGL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1mse_C | 1e-43 | 201 | 303 | 2 | 104 | C-Myb DNA-Binding Domain |
| 1msf_C | 1e-43 | 201 | 303 | 2 | 104 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 180 | 197 | QKRCNRMHVRKGRKLPKR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00385 | DAP | Transfer from AT3G27785 | Download |
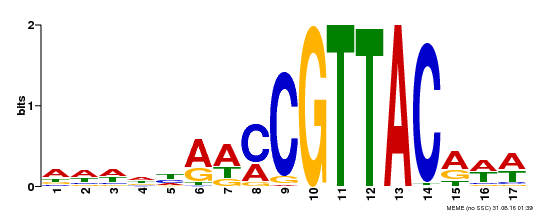
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006446542.1 | 0.0 | transcription factor MYB118 | ||||
| TrEMBL | V4U1M7 | 0.0 | V4U1M7_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006446542.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6052 | 26 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G27785.1 | 6e-76 | myb domain protein 118 | ||||



