 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_10058_f_3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 405aa MW: 46139.1 Da PI: 4.9607 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 112.4 | 3e-35 | 14 | 105 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkellek 96
Fl k+ye+++d+++++++sws++++sf+v+++ efa+++Lpk+Fkhsnf+SF+RQLn+YgF+k++ e+ weF+++ F +g+ +ll++
Neem_10058_f_3 14 FLAKTYEMVDDPSTDSVVSWSASNKSFIVWNPPEFARDLLPKFFKHSNFSSFIRQLNTYGFRKIDPEQ---------WEFANEDFVRGQPHLLKN 99
9********************999****************************************9998.........****************** PP
XXXXXX CS
HSF_DNA-bind 97 ikrkks 102
i+r+k
Neem_10058_f_3 100 IHRRKP 105
***985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46785 | 2.04E-36 | 9 | 103 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 4.0E-64 | 10 | 103 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene3D | G3DSA:1.10.10.10 | 5.6E-39 | 10 | 103 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF00447 | 6.1E-32 | 14 | 103 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.4E-19 | 14 | 37 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.4E-19 | 52 | 64 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 53 | 77 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.4E-19 | 65 | 77 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 405 aa Download sequence Send to blast |
MDEAQGGSNS LPPFLAKTYE MVDDPSTDSV VSWSASNKSF IVWNPPEFAR DLLPKFFKHS 60 NFSSFIRQLN TYGFRKIDPE QWEFANEDFV RGQPHLLKNI HRRKPVHSHS NQNLHGQGNP 120 LTEFERQTLK DDIERLKKEK EMLLLELQKH EQERQGFESQ MQLLKDRFQN MEQRQQRILS 180 LVARALQRQG LASNLGAQLE THDRKRRLPR IGYFYDEANI EDNQTGTSQA VPRVSAETTN 240 VLSSNMEQVE QLESALTFWE NIVQDVGQSC FQPTSSPEFD ESTSCADSPA ISCIQLNVEV 300 RPKSPGIDMN SEPAVTAAPE PVASQEPVAA TTAPAPTGVN DMFWEQFLTE NPGSTDTQEV 360 QSERRDSDGR KSENKTADHG RFWWNMRNVN SLSEQMGHLT PAERT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 8e-25 | 7 | 103 | 22 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 8e-25 | 7 | 103 | 22 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 8e-25 | 7 | 103 | 22 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 202 | 207 | DRKRRL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds DNA sequence 5'-AGAAnnTTCT-3' known as heat shock promoter elements (HSE). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
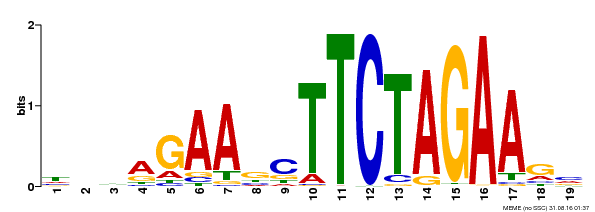
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006449558.1 | 0.0 | heat stress transcription factor A-4a | ||||
| Refseq | XP_024047370.1 | 0.0 | heat stress transcription factor A-4a | ||||
| Refseq | XP_024047371.1 | 0.0 | heat stress transcription factor A-4a | ||||
| Refseq | XP_024047372.1 | 0.0 | heat stress transcription factor A-4a | ||||
| Swissprot | O49403 | 1e-138 | HFA4A_ARATH; Heat stress transcription factor A-4a | ||||
| TrEMBL | V4TQJ7 | 0.0 | V4TQJ7_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006449558.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2853 | 28 | 68 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 1e-125 | heat shock transcription factor A4A | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



