 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araha.5856s0005.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 613aa MW: 66970.1 Da PI: 5.2292 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 26.1 | 1.5e-08 | 256 | 276 | 5 | 26 |
--SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaHg 26
+C+ f++ G C++Gd+C +aHg
Araha.5856s0005.1.p 256 PCPEFRK-GSCSRGDTCEYAHG 276
8******.*************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF12796 | 3.0E-9 | 53 | 123 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 4.97E-13 | 56 | 134 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 13.973 | 57 | 132 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.0044 | 57 | 87 | IPR002110 | Ankyrin repeat |
| Gene3D | G3DSA:1.25.40.20 | 2.1E-13 | 57 | 132 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.62E-13 | 59 | 165 | No hit | No description |
| PROSITE profile | PS50088 | 9.564 | 92 | 127 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.021 | 92 | 124 | IPR002110 | Ankyrin repeat |
| SMART | SM00356 | 4.9E-5 | 251 | 277 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 4.6E-4 | 255 | 278 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 5.4E-6 | 256 | 276 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 13.263 | 256 | 278 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 5.4E-4 | 284 | 314 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 6.221 | 286 | 310 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 20 | 286 | 309 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 613 aa Download sequence Send to blast |
MGVDELSHLK FSLLLESSAC NDLSGFKSLV EEEGLESIDG SGLWYGRRLG SKKMGFEERT 60 PLMIAALFGS KEVVDYIIST GLVDVNRSCG SDGATALHCA VSGLSANSFE IVTLLLKGSA 120 DPDSCDAYGN KPGDMIFPCL SPVFSARMKV LERLLKGNDD LNEVNGQEEG EREVEVEVEV 180 SVSPPRGSER KEYPVDPTLP DIKNGIYGTD EFRMYAFKIK PCSRAYSHDW TECPFVHPGE 240 NARRRDPRKY HYSCVPCPEF RKGSCSRGDT CEYAHGIFEC WLHPAQYRTR LCKDETNCSR 300 RVCFFAHKPE ELRPLYPSTG SGVPSPRSSF SSCNSSSAFD MGPISPLPIG ASTTPPLSPN 360 GVSSPMGGGK TWMNWPNITP PALQLPGSRL KSALNAREID FSEEMQSLTS PTTWNNTPMS 420 AASSPFSGKG MNRLAGGAMS PVNSLSDMFG TEDNTSGLQI RRSVINPQLH SNSLSSSPVG 480 ANSLFSMDSS TVLASRAAEF AKQRSQSFIE RSNGLNHHPA ISSMTTTCLN DWGSLDGKLD 540 WSVQGDELQK LRKSTSFRLR AGGMESRLTN EGTGLEEPDV SWVEPLVKEP QETRLAPVWM 600 EQSYMETEQT VA* |
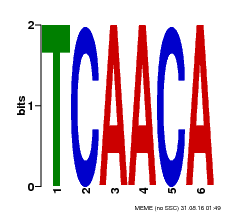
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00566 | DAP | Transfer from AT5G58620 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araha.5856s0005.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB020755 | 0.0 | AB020755.1 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MZN1. | |||
| GenBank | AY054176 | 0.0 | AY054176.1 Arabidopsis thaliana AT5g58620/mzn1_70 mRNA, complete cds. | |||
| GenBank | AY103314 | 0.0 | AY103314.1 Arabidopsis thaliana AT5g58620/mzn1_70 mRNA, complete cds. | |||
| GenBank | CP002688 | 0.0 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020887576.1 | 0.0 | zinc finger CCCH domain-containing protein 66 | ||||
| Refseq | XP_020887584.1 | 0.0 | zinc finger CCCH domain-containing protein 66 | ||||
| Swissprot | Q9LUZ4 | 0.0 | C3H66_ARATH; Zinc finger CCCH domain-containing protein 66 | ||||
| TrEMBL | D7MQF9 | 0.0 | D7MQF9_ARALL; Zinc finger (CCCH-type) family protein | ||||
| STRING | fgenesh2_kg.8__1801__AT5G58620.1 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM877 | 26 | 107 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G58620.1 | 0.0 | C3H family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Araha.5856s0005.1.p |



