 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araha.33550s0002.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 604aa MW: 67141.3 Da PI: 5.8993 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.5 | 5.6e-19 | 59 | 113 | 2 | 56 |
T--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 2 rkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k +++t++q++eLe++F+++++p++++r eL kkl L+ +q+k+WFqNrR+++k
Araha.33550s0002.1.p 59 TKYHRHTSYQIQELESFFKECPHPNEKQRLELGKKLTLESKQIKFWFQNRRTQMK 113
678899**********************************************999 PP
| |||||||
| 2 | START | 153.3 | 2e-48 | 225 | 426 | 3 | 206 |
HHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECT CS
START 3 aeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetla....kaetleviss 87
a ea++el+k+a+++ +W+ ++e++ + +r++g+v ++ lve+l+d++ +W e+++ +t+evis+
Araha.33550s0002.1.p 225 AMEAMDELLKLAELDNLLWSAKI----EKESMNHL--------AGSRETGLVLINSLALVETLMDTN-KWAEMFEcivaVGSTVEVISN 300
56788888888888888886655....66666666........569*********************.**********9********** PP
T......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE- CS
START 88 g......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwveh 169
g g lqlm+ae+q++splvp + f+Ry++q+g+g w++vdvS d +++++ +s+ +++++pSg++i++ +ng skvtw+e
Araha.33550s0002.1.p 301 GtdgsrsGSLQLMQAEFQVMSPLVPiKQKKFLRYCKQHGDGLWAVVDVSYDINRENEYLKSYGCSKKFPSGCIIQDIGNGCSKVTWIEY 389
********************************************************99******************************* PP
EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 170 vdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
+++++++h+l+++l++s++ ga +w+atlqrqce+
Araha.33550s0002.1.p 390 SEYEESHIHSLYQPLLSSSVGLGATKWLATLQRQCES 426
***********************************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 6.0E-20 | 44 | 115 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.21E-18 | 44 | 115 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 5.2E-14 | 53 | 119 | IPR001356 | Homeobox domain |
| PROSITE profile | PS50071 | 16.973 | 55 | 115 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 9.4E-17 | 59 | 113 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 8.58E-16 | 62 | 115 | No hit | No description |
| PROSITE profile | PS50848 | 39.207 | 214 | 429 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.79E-28 | 220 | 427 | No hit | No description |
| CDD | cd08875 | 4.36E-99 | 220 | 425 | No hit | No description |
| SMART | SM00234 | 3.8E-35 | 223 | 426 | IPR002913 | START domain |
| Pfam | PF01852 | 4.4E-42 | 225 | 426 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 8.06E-7 | 486 | 595 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 604 aa Download sequence Send to blast |
MNGDLDVDMS RGDFNPSYFL GKLKDDEFES RSLSDDSFDA LSGDEDKQEQ RPKKKKRKTK 60 YHRHTSYQIQ ELESFFKECP HPNEKQRLEL GKKLTLESKQ IKFWFQNRRT QMKTQLERHE 120 NVILKQENEK LRLENSFLKE SMRGSLCIDC GGAVIPGEVS FEQHQLRIEN AKLKDELDRI 180 CALANRFIGG SISLEQPSNG GIGSQHLPIG HSVSGGTSLM FMDLAMEAMD ELLKLAELDN 240 LLWSAKIEKE SMNHLAGSRE TGLVLINSLA LVETLMDTNK WAEMFECIVA VGSTVEVISN 300 GTDGSRSGSL QLMQAEFQVM SPLVPIKQKK FLRYCKQHGD GLWAVVDVSY DINRENEYLK 360 SYGCSKKFPS GCIIQDIGNG CSKVTWIEYS EYEESHIHSL YQPLLSSSVG LGATKWLATL 420 QRQCESFTTL FSSQDHTGLS LAGTKSILKL AQRMKLNFYS GITASSIHKW EKLLAENGKD 480 QNESSMLILQ ETWNDASGAL VVYAPVDIPS MNVVMSGGDS AYVALLPSGF SILPDGSSSS 540 SDQFDTDGGL VNHESKGCLL TVGFQILVNS LPTAKLNVES VETVNNLIAC TIHKIRAALR 600 IPA* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 49 | 57 | QRPKKKKRK |
| 2 | 52 | 57 | KKKKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds to the DNA sequence 5'-GCATTAAATGC-3'. {ECO:0000269|PubMed:16778018}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00559 | DAP | Transfer from AT5G52170 | Download |
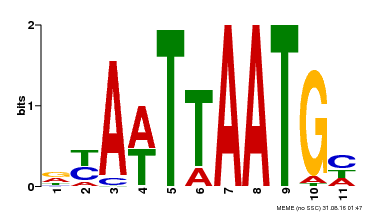
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araha.33550s0002.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB025603 | 0.0 | AB025603.1 Arabidopsis thaliana genomic DNA, chromosome 5, BAC clone:F17P19. | |||
| GenBank | CP002688 | 0.0 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020871109.1 | 0.0 | homeobox-leucine zipper protein HDG7 | ||||
| Swissprot | Q9LTK3 | 0.0 | HDG7_ARATH; Homeobox-leucine zipper protein HDG7 | ||||
| TrEMBL | A0A178URB7 | 0.0 | A0A178URB7_ARATH; HDG7 | ||||
| STRING | AT5G52170.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52170.1 | 0.0 | homeodomain GLABROUS 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Araha.33550s0002.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



