 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aqcoe6G111000.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Ranunculales; Ranunculaceae; Thalictroideae; Aquilegia
|
||||||||
| Family | BES1 | ||||||||
| Protein Properties | Length: 314aa MW: 34474.2 Da PI: 9.6658 | ||||||||
| Description | BES1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF822 | 216 | 9.1e-67 | 5 | 150 | 1 | 150 |
DUF822 1 ggsgrkptwkErEnnkrRERrRRaiaakiyaGLRaqGnyklpkraDnneVlkALcreAGwvvedDGttyrkgskpleeaeaagssasaspes 92
g+sgr+ptwkErEnnkrRERrRRaiaa iy+GLRa Gnyklpk++DnneVlkALc+eAGw+ve+DGttyrkg++p+++ ++g+s+++sp+
Aqcoe6G111000.1.p 5 GTSGRMPTWKERENNKRRERRRRAIAANIYSGLRASGNYKLPKHCDNNEVLKALCAEAGWIVEEDGTTYRKGFRPQSNDTTGGNSTNMSPCL 96
589***************************************************************************************** PP
DUF822 93 slqsslkssalaspvesysaspksssfpspssldsislasaasllpvlsvlslvsssl 150
s q s++ss+++spv+sy+asp+ss+f sps++ds+++ +lp+l++l++v+ssl
Aqcoe6G111000.1.p 97 SHQPSAQSSYFPSPVPSYHASPSSSAFASPSRFDSNPA----YILPFLQNLASVPSSL 150
***********************************976....69*******9999986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05687 | 1.3E-60 | 7 | 131 | IPR008540 | BES1/BZR1 plant transcription factor, N-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009742 | Biological Process | brassinosteroid mediated signaling pathway | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048316 | Biological Process | seed development | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 314 aa Download sequence Send to blast |
MTSRGTSGRM PTWKERENNK RRERRRRAIA ANIYSGLRAS GNYKLPKHCD NNEVLKALCA 60 EAGWIVEEDG TTYRKGFRPQ SNDTTGGNST NMSPCLSHQP SAQSSYFPSP VPSYHASPSS 120 SAFASPSRFD SNPAYILPFL QNLASVPSSL PPIRISNSAP VTPPLPSPNS RSPKRKPDWQ 180 ALSNYNMASY RHPLFIASAP TSPTHRHRHF TPSTIFEGDE SDSSTVDLPQ WASFQTVTTG 240 GAPVSPTYNL VNPVSEQNSP KSGIAGSRGK REHGAEFEFE NRRVKPWEGE MIHEVGVDDL 300 ELTLGNGKTR EFG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zd4_A | 1e-24 | 9 | 89 | 372 | 452 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_B | 1e-24 | 9 | 89 | 372 | 452 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_C | 1e-24 | 9 | 89 | 372 | 452 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_D | 1e-24 | 9 | 89 | 372 | 452 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00237 | DAP | Transfer from AT1G75080 | Download |
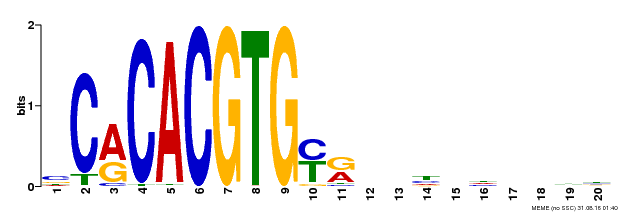
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_023881253.1 | 1e-127 | BES1/BZR1 homolog protein 2-like | ||||
| Swissprot | Q94A43 | 2e-99 | BEH2_ARATH; BES1/BZR1 homolog protein 2 | ||||
| TrEMBL | A0A2G5E9V5 | 0.0 | A0A2G5E9V5_AQUCA; Uncharacterized protein | ||||
| STRING | Aquca_010_00422.1 | 0.0 | (Aquilegia coerulea) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G75080.2 | 3e-80 | BES1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aqcoe6G111000.1.p |



