 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aqcoe5G287400.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Ranunculales; Ranunculaceae; Thalictroideae; Aquilegia
|
||||||||
| Family | M-type_MADS | ||||||||
| Protein Properties | Length: 250aa MW: 28919.9 Da PI: 9.6172 | ||||||||
| Description | M-type_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 47.3 | 2.6e-15 | 11 | 50 | 3 | 42 |
--SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-T CS
SRF-TF 3 ienksnrqvtfskRrngilKKAeELSvLCdaevaviifss 42
i n+s + t+ kR++g++KK +ELS+LC++e++ +++++
Aqcoe5G287400.1.p 11 IANDSAQRSTYKKRKQGLMKKINELSTLCGVEACAVVYGP 50
8899999*******************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 15.558 | 1 | 49 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 1.2E-18 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00266 | 7.96E-32 | 2 | 86 | No hit | No description |
| PRINTS | PR00404 | 2.3E-6 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 1.31E-22 | 3 | 95 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 1.2E-15 | 11 | 50 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.3E-6 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0043078 | Cellular Component | polar nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 250 aa Download sequence Send to blast |
MARKKVKLAW IANDSAQRST YKKRKQGLMK KINELSTLCG VEACAVVYGP YDPQVPDVWP 60 SPSDAHRVLT QFKSLPEMER NKKMMNQEAF LKERMAKMRE QIKKQQRENR EFEITQLMNR 120 TLIDGTGQIL QNVETKELKD LAWMIDEKMK RIQKRIDSLR STMESAVQQI NGTGIQTTTE 180 MEAAQRQIWF MNQIMNNPPP QQEQQQQPTD YYGNNPPTQQ EHHHQPIGYY RGVGNDFANG 240 GMPYADNTP* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 2 | 24 | RKKVKLAWIANDSAQRSTYKKRK |
| 2 | 16 | 24 | QRSTYKKRK |
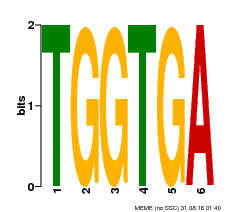
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00553 | DAP | Transfer from AT5G48670 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX680240 | 0.0 | JX680240.1 Aquilegia coerulea MADS-box protein AGL83 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010271057.1 | 8e-68 | PREDICTED: agamous-like MADS-box protein AGL80 | ||||
| TrEMBL | A0A2G5D2X9 | 0.0 | A0A2G5D2X9_AQUCA; Uncharacterized protein | ||||
| STRING | Aquca_029_00010.1 | 0.0 | (Aquilegia coerulea) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G48670.1 | 2e-48 | AGAMOUS-like 80 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aqcoe5G287400.1.p |



