 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco013228.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 697aa MW: 76179.1 Da PI: 5.8777 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.7 | 4.1e-16 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE+ ++++a k++G W +I +++g ++t+ q++s+ qk+
Aco013228.1 24 RERWTEEEHNRFLEALKLYGRA-WQRIEEHIG-TKTAVQIRSHAQKF 68
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 7.48E-17 | 18 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.173 | 19 | 73 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 6.3E-17 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.1E-12 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.8E-13 | 24 | 67 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-8 | 24 | 64 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.29E-9 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010243 | Biological Process | response to organonitrogen compound | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043496 | Biological Process | regulation of protein homodimerization activity | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 697 aa Download sequence Send to blast |
MEANSSGEEF IVKVRKPYTI TKQRERWTEE EHNRFLEALK LYGRAWQRIE EHIGTKTAVQ 60 IRSHAQKFFT KLEKEAMTKG VPPGQAHDID IPPPRPKRKP NSPYPRKSCA LSPLSSTGEV 120 VNEKSNYSFS PLSAGNREVS QKESDAPSEP QNKEMSENGS CSEVLNLFRE APCVSMSSIN 180 KGSAYNHSAY KGFDPMMKES KERSLLEKTP VLTETNEKPD AGTDQETERL KGIRISSDAN 240 CPKNEGVNDQ GSPPVETIQV DQINNPMAEQ AGGAHGESVN PSTNHSLSTG PKLHENSTMP 300 SFHHNYPIFS PFPQCHNQQD AYRSYLNISS TFSNLLVSTL LQNPAIHTVA RFAASFWPSP 360 EINASVDSNP ETLASQNLAS LATATVAAAS AWWATQGLLP LFPPTPLGFA FPPPPTTTGL 420 PIVGDTRTSE KESGDGIYQG PSKDQEAPNL DQSEGLKQYP SSKPSLSSLS DSDESGREEE 480 KRSLCAELKA SRTSKSKPSS ANEGTNNADS LRNNKKQDRS SCGSNTPSSS DVETDMPEND 540 EQINDNSKQD YFGNSLAGDT NYRRFRSSGS TIDSWKEVSE EGRLAFQALF RRGVLPQSFS 600 PPPAEEEGVK CKGGEETTLP PVNNTGLTMD TLIRSNESGE LGSLLTSGFL QAKLKARRTG 660 FKPYKRCSAE AKESRATAGE EAINNKRIRL EGETST* |
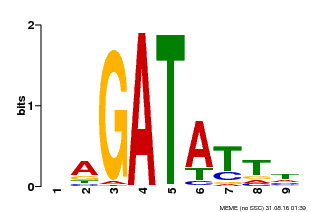
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY750151 | 0.0 | AY750151.1 Ananas comosus circadian clock-associated protein (CCA1) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020114596.1 | 0.0 | protein LHY-like isoform X3 | ||||
| TrEMBL | A0A199VZX4 | 0.0 | A0A199VZX4_ANACO; Protein LHY | ||||
| STRING | OS08T0157600-01 | 1e-180 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 3e-50 | circadian clock associated 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco013228.1 |



