 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco010320.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 380aa MW: 43843.8 Da PI: 8.6026 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 18.6 | 5.1e-06 | 79 | 100 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg sF + +Lk+H+++H
Aco010320.1 79 SCEECGASFQKPAHLKQHMQSH 100
6*******************99 PP
| |||||||
| 2 | zf-C2H2 | 19.6 | 2.5e-06 | 106 | 130 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++Cp dC +s++rk++L+rH+ tH
Aco010320.1 106 FSCPveDCRQSYRRKDHLTRHLHTH 130
89********************999 PP
| |||||||
| 3 | zf-C2H2 | 13.2 | 0.00027 | 135 | 160 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+ C+ C+++F+ k n++rH + H
Aco010320.1 135 FACTieGCDRRFNIKANMQRHVKErH 160
679999****************9988 PP
| |||||||
| 4 | zf-C2H2 | 21.9 | 4.4e-07 | 172 | 194 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y C+ Cgk+F+ s L++H +H
Aco010320.1 172 YACKECGKTFKYASKLRKHEDSH 194
89*****************9888 PP
| |||||||
| 5 | zf-C2H2 | 17.8 | 9.1e-06 | 261 | 285 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
+C+ dC+ sFs++snL++Hi+ H
Aco010320.1 261 QCSfkDCHLSFSNRSNLNKHIKAcH 285
69999*****************955 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 18 | 31 | 53 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.921 | 78 | 105 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0088 | 78 | 100 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 7.2E-5 | 80 | 96 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 80 | 100 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.8E-10 | 90 | 132 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-13 | 97 | 132 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.012 | 106 | 130 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.741 | 106 | 135 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 108 | 130 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.24E-7 | 133 | 194 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.7E-8 | 133 | 161 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.803 | 135 | 165 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0012 | 135 | 160 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 137 | 160 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 4.8E-7 | 169 | 194 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0025 | 172 | 194 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.445 | 172 | 194 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 174 | 194 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3 | 202 | 227 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 204 | 227 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 24 | 230 | 251 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 3.83E-7 | 241 | 297 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.9E-6 | 241 | 282 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.344 | 260 | 290 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0058 | 260 | 285 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 262 | 285 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 2.7 | 291 | 317 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 293 | 317 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 380 aa Download sequence Send to blast |
MACLESDASE GEVESEGCAK KQPFFRDIRR YSCEFCGIIR SKKSLIRSHI LAHHKDEYKG 60 SQICEDSEEN MARKDVSRSC EECGASFQKP AHLKQHMQSH SLERPFSCPV EDCRQSYRRK 120 DHLTRHLHTH EGKLFACTIE GCDRRFNIKA NMQRHVKERH EDGSPCENKK QYACKECGKT 180 FKYASKLRKH EDSHVKLDCV EVVCCEPGCF KTFTNAECLK AHVQFCHQHT QCEICGTKQM 240 KRNIKRHKRK HEGVIVAERV QCSFKDCHLS FSNRSNLNKH IKACHEQLKP FVCRLSGCGQ 300 KFPYRHVRDN HEKSGSHVYL QGDFLEVDEQ SQSRCKGGRK RKTLSVETLT RKRVAPPGEA 360 SCLDDGTEYM RWLLSGDTQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2i13_A | 2e-17 | 79 | 253 | 23 | 186 | Aart |
| 2i13_B | 2e-17 | 79 | 253 | 23 | 186 | Aart |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
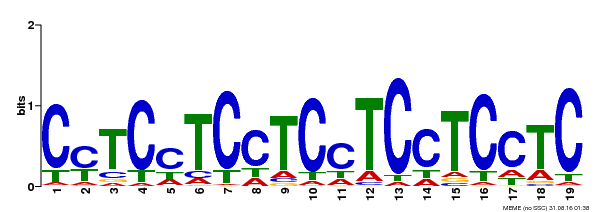
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020079780.1 | 0.0 | transcription factor IIIA isoform X2 | ||||
| Swissprot | Q84MZ4 | 1e-118 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A199UVM7 | 0.0 | A0A199UVM7_ANACO; Transcription factor IIIA | ||||
| STRING | XP_008793643.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-120 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco010320.1 |



